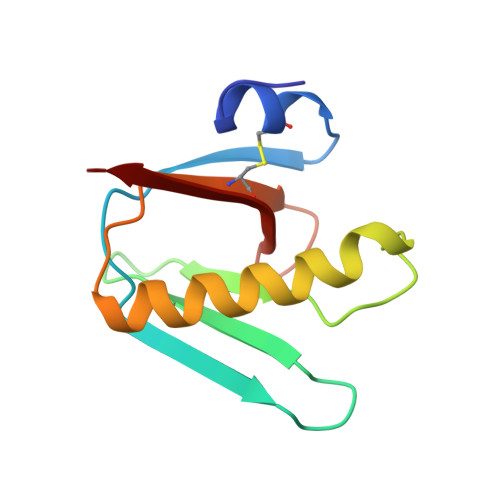
Crystal structure of heat-labile enterotoxin from Escherichia coli with increased thermostability introduced by an engineered disulfide bond in the A subunit.
van den Akker, F., Feil, I.K., Roach, C., Platas, A.A., Merritt, E.A., Hol, W.G.(1997) Protein Sci 6: 2644-2649
- PubMed: 9416616
- DOI: https://doi.org/10.1002/pro.5560061219
- Primary Citation Related Structures:
1LT3 - PubMed Abstract:
Cholera toxin (CT) produced by Vibrio cholerae and heat-labile enterotoxin (LT-I), produced by enterotoxigenic Escherichia coli, are AB5 heterohexamers with an ADP-ribosylating A subunit and a GM1 receptor binding B pentamer. These toxins are among the most potent mucosal adjuvants known and, hence, are of interest both for the development of anti-diarrheal vaccines against cholera or enterotoxigenic Escherichia coli diarrhea and also for vaccines in general. However, the A subunits of CT and LT-I are known to be relatively temperature sensitive. To improve the thermostability of LT-I an additional disulfide bond was introduced in the A1 subunit by means of the double mutation N40C and G166C. The crystal structure of this double mutant of LT-I has been determined to 2.0 A resolution. The protein structure of the N40C/G166C double mutant is very similar to the native structure except for a few local shifts near the new disulfide bond. The introduction of this additional disulfide bond increases the thermal stability of the A subunit of LT-I by 6 degrees C. The enhancement in thermostability could make this disulfide bond variant of LT-I of considerable interest for the design of enterotoxin-based vaccines.
- Department of Biochemistry, University of Washington, Seattle 98195-7420, USA.
Organizational Affiliation: