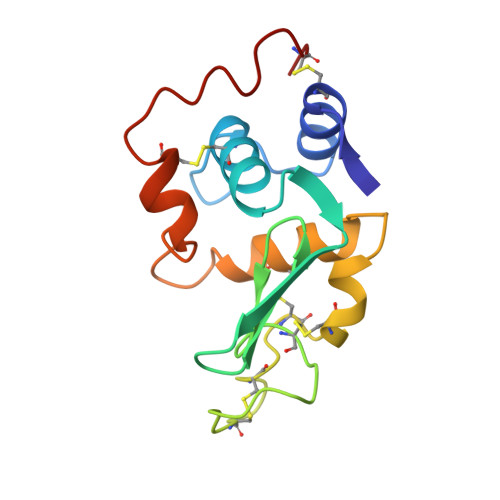
Structure of a conformationally constrained Arg-Gly-Asp sequence inserted into human lysozyme.
Yamada, T., Song, H., Inaka, K., Shimada, Y., Kikuchi, M., Matsushima, M.(1995) J Biological Chem 270: 5687-5690
- PubMed: 7890692 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.270.11.5687
- Primary Citation Related Structures:
1LMT - PubMed Abstract:
To examine the effect of a conformational constraint introduced into the Arg-Gly-Asp (RGD) sequence on cell adhesion activity, we constructed a mutant protein by inserting an RGD-containing sequence flanked by two Cys residues between Val74 and Asn75 of human lysozyme. The CRGDSC-inserted lysozyme was expressed in yeast, purified, and designated as Cys-RGD4. Using baby hamster kidney cells, Cys-RGD4 was shown to possess even higher cell adhesion activity than that of the RGDS-inserted lysozyme, RGD4. The Cys-RGD4 protein was co-crystallized with a lysozyme inhibitor, tri-N-acetylchitotriose, and the three-dimensional structure was determined at 1.6-A resolution by x-ray crystallography. In contrast to RGD4, the inserted RGD-containing region of Cys-RGD4 was well defined. The structural analysis revealed that the two inserted Cys residues form a new disulfide bond in Cys-RGD4, as expected, and that the RGD region assumes a type II' beta-turn conformation of Gly-Asp with a hydrogen bond between the C = O of Arg and the H-N of Ser. In addition, it was confirmed that two more hydrogen bonds are present in the RGD region of the Cys-RGD4 lysozyme. These results suggest that the conformation of the RGD-containing region is rigid and stable in the Cys-RGD4 molecule and that the type II' beta-turn structure of RGD is essential for binding to integrins with high affinity.
- Protein Engineering Research Institute, Osaka, Japan.
Organizational Affiliation: