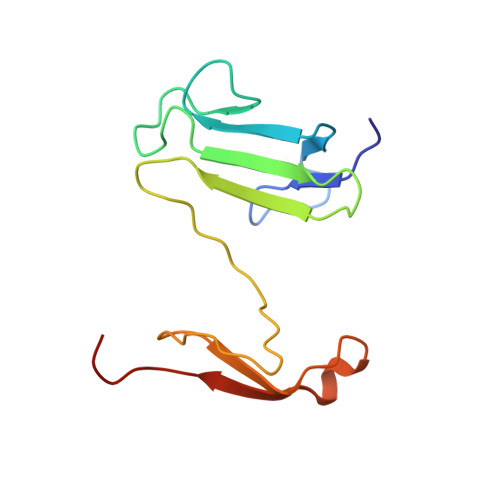
Crystal structure of the FHA domain of the Chfr mitotic checkpoint protein and its complex with tungstate.
Stavridi, E.S., Huyen, Y., Loreto, I.R., Scolnick, D.M., Halazonetis, T.D., Pavletich, N.P., Jeffrey, P.D.(2002) Structure 10: 891-899
- PubMed: 12121644 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(02)00776-1
- Primary Citation Related Structures:
1LGP, 1LGQ - PubMed Abstract:
The Chfr mitotic checkpoint protein is frequently inactivated in human cancer. We determined the three-dimensional structure of its FHA domain in its native form and in complex with tungstate, an analog of phosphate. The structures revealed a beta sandwich fold similar to the previously determined folds of the Rad53 N- and C-terminal FHA domains, except that the Rad53 domains were monomeric, whereas the Chfr FHA domain crystallized as a segment-swapped dimer. The ability of the Chfr FHA domain to recognize tungstate suggests that it shares the ability with other FHA domains to bind phosphoproteins. Nevertheless, differences in the sequence and structure of the Chfr and Rad53 FHA domains suggest that FHA domains can be divided into families with distinct binding properties.
- Molecular Genetics Program, Structural Biology Program, Wistar Institute, Philadelphia, PA 19104, USA.
Organizational Affiliation: