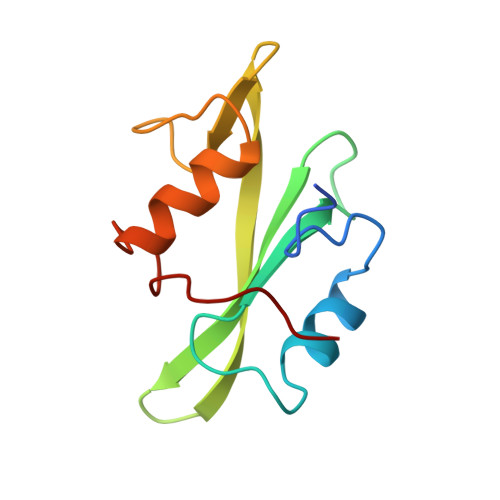
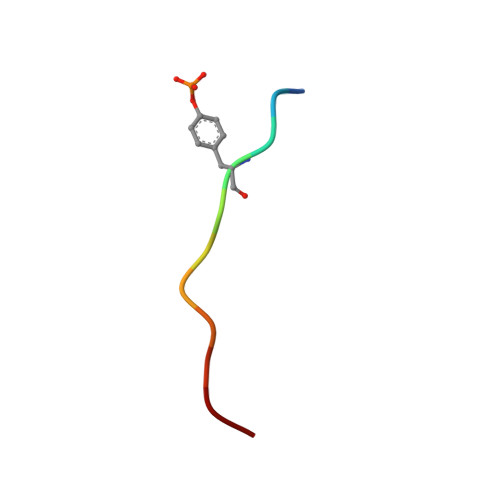
Recognition of a high-affinity phosphotyrosyl peptide by the Src homology-2 domain of p56lck.
Eck, M.J., Shoelson, S.E., Harrison, S.C.(1993) Nature 362: 87-91
- PubMed: 7680435
- DOI: https://doi.org/10.1038/362087a0
- Primary Citation of Related Structures:
1LCJ - PubMed Abstract:
The Src homology-2 (SH2) domains are modules of about 100 amino-acid residues that are found in many intracellular signal-transduction proteins. They bind phosphotyrosine-containing sequences with high affinity and specificity, recognizing phosphotyrosine in the context of the immediately adjacent polypeptide sequence. The protein p56lck (Lck) is a Src-like, lymphocyte-specific tyrosine kinase. A phosphopeptide library screen has recently been used to deduce an 'optimal' binding sequence for the Lck SH2 domain. There is selectivity for the residues Glu, Glu and Ile in the three positions C-terminal to the phosphotyrosine. An 11-residue phosphopeptide derived from the hamster polyoma middle-T antigen, EPQpYEEIPIYL, binds with an approximately 1 nM dissociation constant to the Lck SH2 (ref. 17), an affinity equivalent to that of the tightest known SH2-phosphopeptide complex. We report here the high-resolution crystallographic analysis of the Lck SH2 domain in complex with this phosphopeptide. Recent crystallographically derived structures of the Src SH2 domain in complex with low-affinity peptides, which do not contain the EEI consensus, and NMR-derived structures of unliganded Abl (ref. 19) and p85 (ref. 20) SH2 domains have revealed the conserved fold of the SH2 domain and the properties of a phosphotyrosine binding pocket. Our high-affinity complex shows the presence of a second pocket for the residue (pY + 3) three positions C-terminal to the phosphotyrosine (pY). The peptide is anchored by insertion of the pY and pY + 3 side chains into their pockets and by a network of hydrogen bonds to the peptide main chain. In the low-affinity phosphopeptide/Src complexes, the pY + 3 residues do not insert into the homologous binding pocket and the peptide main chain remains displaced from the surface of the domain.
- Howard Hughes Medical Institute, Children's Hospital, Boston, Massachusetts 02115.
Organizational Affiliation: