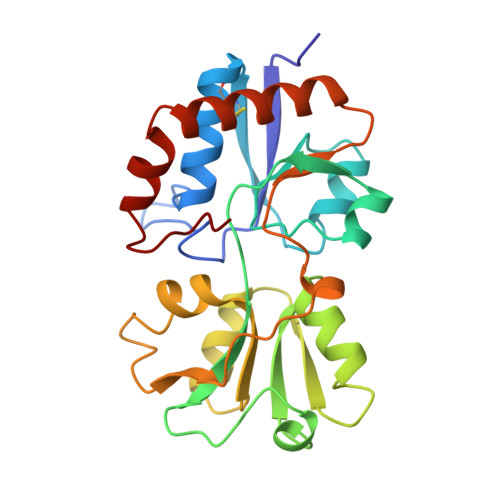
Structural basis for multiple ligand specificity of the periplasmic lysine-, arginine-, ornithine-binding protein.
Oh, B.H., Ames, G.F., Kim, S.H.(1994) J Biological Chem 269: 26323-26330
- PubMed: 7929349 Search on PubMed
- Primary Citation Related Structures:
1LAF, 1LAG, 1LAH - PubMed Abstract:
The substrate-binding site of a protein with multiple specificity should satisfy geometric and energetic complementarity for several different substrates. The structural basis of the multiple ligand specificity of the periplasmic lysine-, arginine-, ornithine-binding protein (LAO) was investigated by determining and analyzing the structures of the protein unliganded and liganded with each of the three high-affinity ligands (L-lysine, L-arginine, and L-ornithine) and with one low-affinity ligand (L-histidine). The geometric complementarity is achieved primarily by virtue of the large size of the ligand-binding site which can accommodate the maximum common volume of the four ligands plus three water molecules. The optimization of energetic complementarity is achieved by the relocation of protein-bound water molecules and by the movement of the Asp-11 side chain. The structure of the LAO-histidine complex indicates that the 30-fold reduced affinity of the protein for histidine is primarily due to unavailability of one ionic interaction of the histidine side chain with the protein which is present in the other three complexes.
- Department of Chemistry, Lawrence Berkeley Laboratory, Berkeley, California.
Organizational Affiliation: