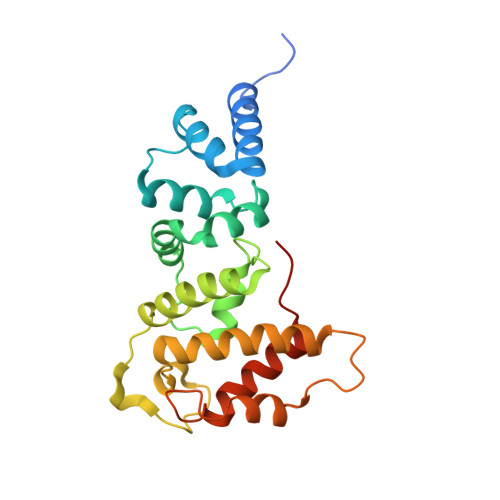
Mechanism of domain closure of Sec7 domains and role in BFA sensitivity.
Renault, L., Christova, P., Guibert, B., Pasqualato, S., Cherfils, J.(2002) Biochemistry 41: 3605-3612
- PubMed: 11888276
- DOI: https://doi.org/10.1021/bi012123h
- Primary Citation of Related Structures:
1KU1 - PubMed Abstract:
Activation of small G proteins of the Arf family is initiated by guanine nucleotide exchange factors whose catalytic Sec7 domain stimulates the dissociation of the tightly bound GDP nucleotide. The exchange reaction involves distinct sequential steps that can be trapped by the noncompetitive inhibitor brefeldin A, by mutation of an invariant catalytic glutamate, or by removal of guanine nucleotides. Arf-GDP retains most characteristics of its GDP-bound form at the initial low-affinity Arf-GDP-Sec7 step. It then undergoes large conformational changes toward its GTP-bound form at the next step, and eventually dissociates GDP to form a nucleotide-free high-affinity Arf-Sec7 complex at the last step. Thus, Arf proteins evolve through different conformations that must be accommodated by Sec7 domains in the course of the reaction. Here the contribution of the flexibility of Sec7 domains to the exchange reaction was investigated with the crystal structure of the unbound Sec7 domain of yeast Gea2. Comparison with Gea2 in complex with nucleotide-free Arf1 Delta 17 [Goldberg, J. (1998) Cell 95, 237-248] reveals that Arf induces closure of the two subdomains that form the sides of its active site. Several residues that determine sensitivity to brefeldin A are involved in interdomain and local movements, pointing to the importance of the flexibility of Sec7 domains for the inhibition mechanism. Altogether, this suggests a model for the initial steps of the exchange reaction where Arf docks onto the C-terminal domain of the Sec7 domain before closure of the N-terminal domain positions the catalytic glutamate to complete the reaction.
- Laboratoire d'Enzymologie et Biochimie Structurales, UPR 9063 CNRS, 1, avenue de la Terrasse, 91198 Gif sur Yvette cedex, France.
Organizational Affiliation: