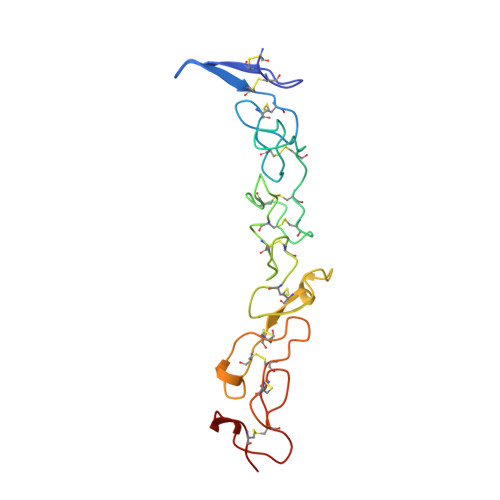
Crystal structure of three consecutive laminin-type epidermal growth factor-like (LE) modules of laminin gamma1 chain harboring the nidogen binding site.
Stetefeld, J., Mayer, U., Timpl, R., Huber, R.(1996) J Mol Biology 257: 644-657
- PubMed: 8648630 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1996.0191
- Primary Citation Related Structures:
1KLO - PubMed Abstract:
The structure of three consecutive laminin-type EGF-like (LE) modules of mouse laminin gammma1 chain, gamma1III3-5 (positions 738 to 899), has been determined by multiple isomorphous replacement in a crystal of space group p6(4)22 (a=b=74.57 angstroms, c = 185.11 angstroms and gamma = 120 degrees). The crystal structure was refined using restrained crystallographic refinement to an R-factor of 19.72% for 14,983 independent reflections with intensities F(obs)> 0 at 2.1 angstroms resolution, with root mean square deviation of 0.012 angstroms and 1.690 degrees from ideal bond lengths and bond angles, respectively. The final model consisted of 1179 (non-hydrogen) protein atoms within 162 residues and 119 water molecules. The molecule showed a rod-like structure of about 76 angstroms length with individual modules twisted relative to each other by about 70 degrees. Each module has the same disulfide bond connections Cys1-Cys3 (loop a), Cys2-Cys4 (loop b), Cys5-Cys6 (loop c) and Cys7-Cys8 (loop d), the first three being identical to epidermal growth factor (EGF). All three LE modules showed little secondary structure which was mainly restricted to loop d, but they differed in several other details of their structure. The interface contacts between the LE modules are based on hydrogen bonds and hydrophobic interactions between the hydrophobic core of loop d of the preceding module and the first cysteine and an exposed residue in loop b of the following module. Module 4 was previously shown to contribute the major nidogen binding site of laminis and site-directed mutagenesis demonstrated a specific binding role for Asp800, Asn802, Val804 and Tyr819 in loops a and c. The side-chain of these four residues are all located on the surface in a linear array and separated by a distance of 17 angstroms between Tyr819 and Val804. The entire nidogen binding site is stabilized via main-chain hydrogen bonds which are in part derived from the link between loops b and c (residues Leu815 and Lys816). The data demonstrate the unique nature of the LE modules and only a remote similarity to EGF. They also indicate that the crucial residues in the binding loops provide direct contacts with nidogen and explain the synergism between loops a and c which is essential for binding.
- Abteilungen für Strukturforschung and Proteinchemie, Max-Planck-Institut für Biochemie, Martinsried, Germany.
Organizational Affiliation: