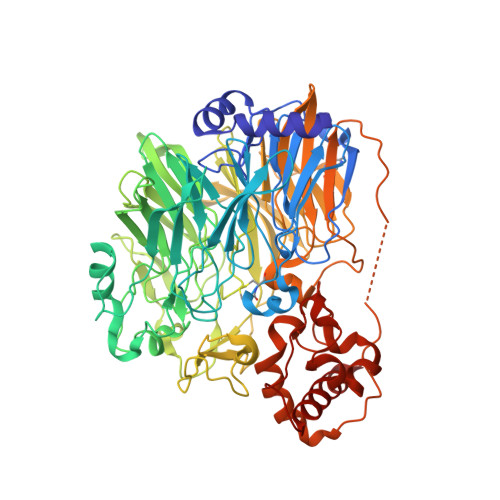
Crystal structure of quinohemoprotein alcohol dehydrogenase from Comamonas testosteroni: structural basis for substrate oxidation and electron transfer.
Oubrie, A., Rozeboom, H.J., Kalk, K.H., Huizinga, E.G., Dijkstra, B.W.(2002) J Biological Chem 277: 3727-3732
- PubMed: 11714714 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M109403200
- Primary Citation Related Structures:
1KB0 - PubMed Abstract:
Quinoprotein alcohol dehydrogenases are redox enzymes that participate in distinctive catabolic pathways that enable bacteria to grow on various alcohols as the sole source of carbon and energy. The x-ray structure of the quinohemoprotein alcohol dehydrogenase from Comamonas testosteroni has been determined at 1.44 A resolution. It comprises two domains. The N-terminal domain has a beta-propeller fold and binds one pyrroloquinoline quinone cofactor and one calcium ion in the active site. A tetrahydrofuran-2-carboxylic acid molecule is present in the substrate-binding cleft. The position of this oxidation product provides valuable information on the amino acid residues involved in the reaction mechanism and their function. The C-terminal domain is an alpha-helical type I cytochrome c with His(608) and Met(647) as heme-iron ligands. This is the first reported structure of an electron transfer system between a quinoprotein alcohol dehydrogenase and cytochrome c. The shortest distance between pyrroloquinoline quinone and heme c is 12.9 A, one of the longest physiological edge-to-edge distances yet determined between two redox centers. A highly unusual disulfide bond between two adjacent cysteines bridges the redox centers. It appears essential for electron transfer. A water channel delineates a possible pathway for proton transfer from the active site to the solvent.
- Laboratory of Biophysical Chemistry and BIOSON Research Institute, University of Groningen, Nijenborgh 4, 9747 AG Groningen, The Netherlands.
Organizational Affiliation: