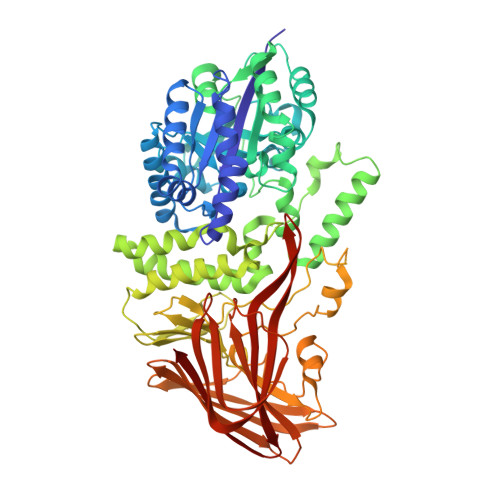
Crystal structures of 4-alpha-glucanotransferase from Thermococcus litoralis and its complex with an inhibitor
Imamura, H., Fushinobu, S., Yamamoto, M., Kumasaka, T., Jeon, B.S., Wakagi, T., Matsuzawa, H.(2003) J Biological Chem 278: 19378-19386
- PubMed: 12618437
- DOI: https://doi.org/10.1074/jbc.M213134200
- Primary Citation of Related Structures:
1K1W, 1K1X, 1K1Y - PubMed Abstract:
Thermococcus litoralis 4-alpha-glucanotransferase (TLGT) belongs to glucoside hydrolase family 57 and catalyzes the disproportionation of amylose and the formation of large cyclic alpha-1,4-glucan (cycloamylose) from linear amylose. We determined the crystal structure of TLGT with and without an inhibitor, acarbose. TLGT is composed of two domains: an N-terminal domain (domain I), which contains a (beta/alpha)7 barrel fold, and a C-terminal domain (domain II), which has a twisted beta-sandwich fold. In the structure of TLGT complexed with acarbose, the inhibitor was bound at the cleft within domain I, indicating that domain I is a catalytic domain of TLGT. The acarbose-bound structure also clarified that Glu123 and Asp214 were the catalytic nucleophile and acid/base catalyst, respectively, and revealed the residues involved in substrate binding. It seemed that TLGT produces large cyclic glucans by preventing the production of small cyclic glucans by steric hindrance, which is achieved by three lids protruding into the active site cleft, as well as an extended active site cleft. Interestingly, domain I of TLGT shares some structural features with the catalytic domain of Golgi alpha-mannosidase from Drosophila melanogaster, which belongs to glucoside hydrolase family 38. Furthermore, the catalytic residue of the two enzymes is located in the same position. These observations suggest that families 57 and 38 evolved from a common ancestor.
- Department of Biotechnology, The University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo 113-8657, Japan. himamur-ra@res.titech.ac.jp
Organizational Affiliation: