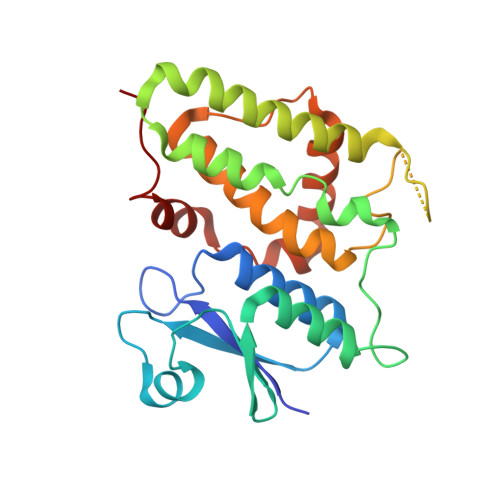
Crystal structure of a soluble form of the intracellular chloride ion channel CLIC1 (NCC27) at 1.4-A resolution.
Harrop, S.J., DeMaere, M.Z., Fairlie, W.D., Reztsova, T., Valenzuela, S.M., Mazzanti, M., Tonini, R., Qiu, M.R., Jankova, L., Warton, K., Bauskin, A.R., Wu, W.M., Pankhurst, S., Campbell, T.J., Breit, S.N., Curmi, P.M.(2001) J Biological Chem 276: 44993-45000
- PubMed: 11551966 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M107804200
- Primary Citation Related Structures:
1K0M, 1K0N, 1K0O - PubMed Abstract:
CLIC1 (NCC27) is a member of the highly conserved class of chloride ion channels that exists in both soluble and integral membrane forms. Purified CLIC1 can integrate into synthetic lipid bilayers forming a chloride channel with similar properties to those observed in vivo. The structure of the soluble form of CLIC1 has been determined at 1.4-A resolution. The protein is monomeric and structurally homologous to the glutathione S-transferase superfamily, and it has a redox-active site resembling glutaredoxin. The structure of the complex of CLIC1 with glutathione shows that glutathione occupies the redox-active site, which is adjacent to an open, elongated slot lined by basic residues. Integration of CLIC1 into the membrane is likely to require a major structural rearrangement, probably of the N-domain (residues 1-90), with the putative transmembrane helix arising from residues in the vicinity of the redox-active site. The structure indicates that CLIC1 is likely to be controlled by redox-dependent processes.
- Initiative for Biomolecular Structure, School of Physics and the Department of Medicine, University of New South Wales, New South Wales 2052, Australia.
Organizational Affiliation: