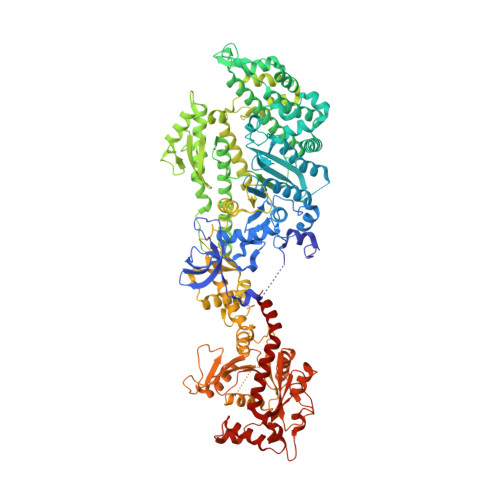
Crystal structure of a dynamin GTPase domain in both nucleotide-free and GDP-bound forms.
Niemann, H.H., Knetsch, M.L., Scherer, A., Manstein, D.J., Kull, F.J.(2001) EMBO J 20: 5813-5821
- PubMed: 11689422 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/20.21.5813
- Primary Citation Related Structures:
1JWY, 1JX2 - PubMed Abstract:
Dynamins form a family of multidomain GTPases involved in endocytosis, vesicle trafficking and maintenance of mitochondrial morphology. In contrast to the classical switch GTPases, a force-generating function has been suggested for dynamins. Here we report the 2.3 A crystal structure of the nucleotide-free and GDP-bound GTPase domain of Dictyostelium discoideum dynamin A. The GTPase domain is the most highly conserved region among dynamins. The globular structure contains the G-protein core fold, which is extended from a six-stranded beta-sheet to an eight-stranded one by a 55 amino acid insertion. This topologically unique insertion distinguishes dynamins from other subfamilies of GTP-binding proteins. An additional N-terminal helix interacts with the C-terminal helix of the GTPase domain, forming a hydrophobic groove, which could be occupied by C-terminal parts of dynamin not present in our construct. The lack of major conformational changes between the nucleotide-free and the GDP-bound state suggests that mechanochemical rearrangements in dynamin occur during GTP binding, GTP hydrolysis or phosphate release and are not linked to loss of GDP.
- Department of Biophysics, Max-Planck-Institute for Medical Research, Jahnstrasse 29, D-69120 Heidelberg, Germany.
Organizational Affiliation: