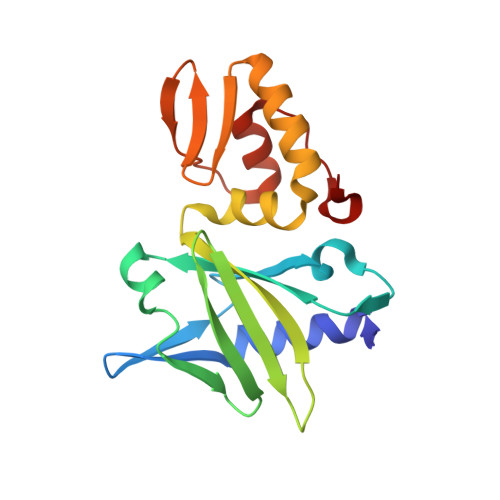
Solution structure of a viral DNA repair polymerase.
Maciejewski, M.W., Shin, R., Pan, B., Marintchev, A., Denninger, A.(2001) Nat Struct Biol 8: 936-941
- PubMed: 11685238
- DOI: https://doi.org/10.1038/nsb1101-936
- Primary Citation of Related Structures:
1JAJ - PubMed Abstract:
DNA polymerase X (Pol X) from the African swine fever virus (ASFV) specifically binds intermediates in the single-nucleotide base-excision repair process, an activity indicative of repair function. In addition, Pol X catalyzes DNA polymerization with low nucleotide-insertion fidelity. The structural mechanisms by which DNA polymerases confer high or low fidelity in DNA polymerization remain to be elucidated. The three-dimensional structure of Pol X has been determined. Unlike other DNA polymerases, Pol X is formed from only a palm and a C-terminal subdomain. Pol X has a novel palm subdomain fold, containing a positively charged helix at the DNA binding surface. Purine deoxynucleoside triphosphate (dNTP) substrates bind between the palm and C-terminal subdomain, at a dNTP-binding helix, and induce a unique conformation in Pol X. The purine dNTP-bound conformation and high binding affinity for dGTP-Mg(2+) of Pol X may contribute to its low fidelity.
- Department of Biochemistry, University of Connecticut Health Center, 263 Farmington Avenue, Farmington, Connecticut 06032, USA.
Organizational Affiliation: