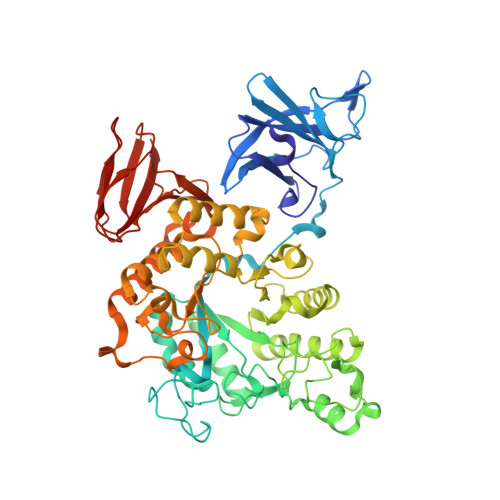
Three-dimensional structure and substrate binding of Bacillus stearothermophilus neopullulanase
Hondoh, H., Kuriki, T., Matsuura, Y.(2003) J Mol Biology 326: 177-188
- PubMed: 12547200
- DOI: https://doi.org/10.1016/s0022-2836(02)01402-x
- Primary Citation Related Structures:
1J0H, 1J0I, 1J0J, 1J0K - PubMed Abstract:
Crystal structures of Bacillus stearothermophilus TRS40 neopullulanase and its complexes with panose, maltotetraose and isopanose were determined at resolutions of 1.9, 2.4, 2.8 and 3.2A, respectively. Since the latter two carbohydrates are substrates of this enzyme, a deactivated mutant at the catalytic residue Glu357-->Gln was used for complex crystallization. The structures were refined at accuracies with r.m.s. deviations of bond lengths and bond angles ranging from 0.005A to 0.008A and 1.3 degrees to 1.4 degrees, respectively. The active enzyme forms a dimer in the crystalline state and in solution. The monomer enzyme is composed of four domains, N, A, B and C, and has a (beta/alpha)(8)-barrel in domain A. The active site lies between domain A and domain N from the other monomer. The results show that dimer formation makes the active-site cleft narrower than those of ordinary alpha-amylases, which may contribute to the unique substrate specificity of this enzyme toward both alpha-1,4 and alpha-1,6-glucosidic linkages. This specificity may be influenced by the subsite structure. Only subsites -1 and -2 are commonly occupied by the product and substrates, suggesting that equivocal recognition occurs at the other subsites, which contributes to the wide substrate specificity of this enzyme.
- Institute for Protein Research, Osaka University, 3-2 Yamada-oka, Suita, 565-0871, Osaka, Japan.
Organizational Affiliation: