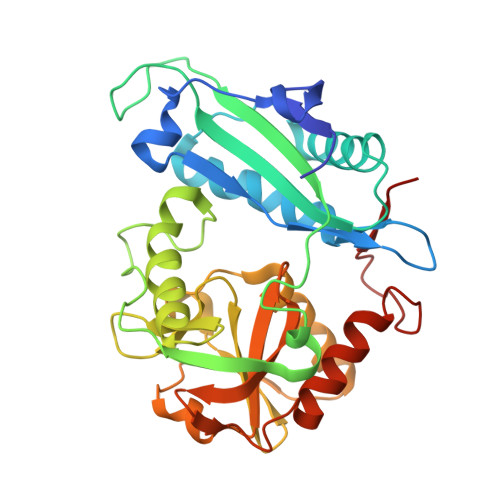
Crystal structures of branched-chain amino Acid aminotransferase complexed with glutamate and glutarate: true reaction intermediate and double substrate recognition of the enzyme.
Goto, M., Miyahara, I., Hayashi, H., Kagamiyama, H., Hirotsu, K.(2003) Biochemistry 42: 3725-3733
- PubMed: 12667063 Search on PubMed
- DOI: https://doi.org/10.1021/bi026722f
- Primary Citation Related Structures:
1IYD, 1IYE - PubMed Abstract:
Branched-chain amino acid aminotransferase (BCAT), which has pyridoxal 5'-phosphate as a cofactor, is a key enzyme in the biosynthetic pathway of hydrophobic amino acids (leucine, isoleucine, and valine). The enzyme reversibly catalyzes the transfer of the amino group of a hydrophobic amino acid to 2-oxoglutarate to form a 2-oxo acid and glutamate. Therefore, the active site of BCAT should have a mechanism to enable recognition of an acidic amino acid as well as a hydrophobic amino acid (double substrate recognition). The three-dimensional structures of Escherichia coli BCAT (eBCAT) in complex with the acidic substrate (glutamate) and the acidic substrate analogue (glutarate) have been determined by X-ray diffraction at 1.82 and 2.15 A resolution, respectively. The enzyme is a homo hexamer, with the polypeptide chain of the subunit folded into small and large domains, and an interdomain loop. The eBCAT in complex with the natural substrate, glutamate, was assigned as a ketimine as the most probable form based upon absorption spectra of the crystal complex and the shape of the residual electron density corresponding to the cofactor-glutamate bond structure. Upon binding of an acidic substrate, the interdomain loop approaches the substrate to shield it from the solvent region, as observed in the complex with a hydrophobic substrate. Both the acidic and the hydrophobic side chains of the substrates are bound to almost the same position in the pocket of the enzyme and are identical in structure. The inner side of the pocket is mostly hydrophobic to accommodate the hydrophobic side chain but has four sites to coordinate with the gamma-carboxylate of glutamate. The mechanism for the double substrate recognition observed in eBCAT is in contrast to those in aromatic amino acid and histidinol-phosphate aminotransferases. In an aromatic amino acid aminotransferase, the acidic side chain is located at the same position as that for the aromatic side chain because of large-scale rearrangements of the hydrogen bond network. In the histidinol-phosphate aminotransferase, the acidic and basic side chains are located at different sites and interact with different residues of the disordered loop.
- Department of Chemistry, Graduate School of Science, Osaka City University, Sugimoto, Sumiyoshi-ku, Osaka 558-8585 Japan.
Organizational Affiliation: