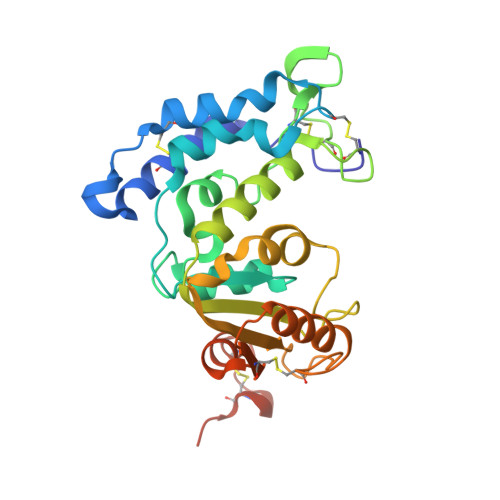
Crystallographic studies on human BST-1/CD157 with ADP-ribosyl cyclase and NAD glycohydrolase activities.
Yamamoto-Katayama, S., Ariyoshi, M., Ishihara, K., Hirano, T., Jingami, H., Morikawa, K.(2002) J Mol Biology 316: 711-723
- PubMed: 11866528 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2001.5386
- Primary Citation Related Structures:
1ISF, 1ISG, 1ISH, 1ISI, 1ISJ, 1ISM - PubMed Abstract:
cADPR is the novel second messenger that elicits calcium release from intracellular calcium stores and works independently of IP(3). In mammals, the ADP-ribosyl cyclase function is found in two membrane proteins, CD38 and BST-1/CD157. These enzymes, exposed extracellularly, bear cADPR hydrolase and NAD glycohydrolase activities. In spite of its functional importance, the structural basis of these enzymatic reactions remains elusive. We determined the crystal structures of the extracellular region of human BST-1 at atomic resolution in the free form and in complexes with five substrate analogues: nicotinamide, NMN, ATPgammaS, ethenoNADP, and ethenoNAD. The three-dimensional structural views of the reaction centre with these ligands revealed the mode of substrate binding and the catalytic mechanism of the multifunctional enzymatic reactions. In each catalytic cleft of the dimeric enzyme, substrates are recognized predominantly through van der Waals interactions with two tryptophan residues, and thereby the N-glycosidic bond of NAD is correctly exposed near a catalytic glutamate residue. Its carboxyl side-chain stabilizes the catalytic intermediate of the S(N)-1 type reaction. This conformation of the catalytic cleft also implies the mechanism of cyclization between the adenine base and the ribose. The three key residues are invariant among the sequences of BST-1, CD38, and Aplysia cyclase, and hence this substrate recognition mode and catalytic scheme appear to be common in the cyclase family.
- Department of Molecular Biology, Research Institute (BERI), Suita-City, Osaka, Japan.
Organizational Affiliation: