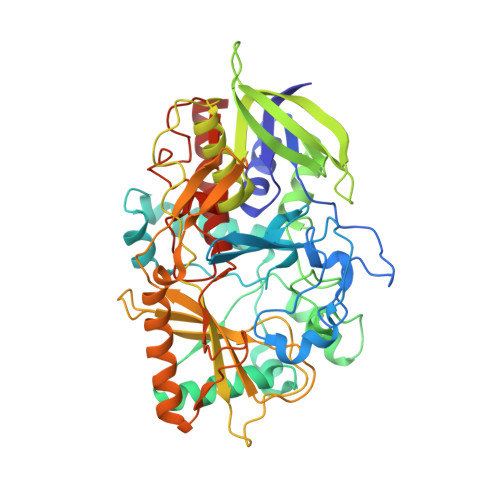
The presence of a hydrogen bond between asparagine 485 and the pi system of FAD modulates the redox potential in the reaction catalyzed by cholesterol oxidase.
Yin, Y., Sampson, N.S., Vrielink, A., Lario, P.I.(2001) Biochemistry 40: 13779-13787
- PubMed: 11705367
- DOI: https://doi.org/10.1021/bi010843i
- Primary Citation of Related Structures:
1IJH - PubMed Abstract:
Cholesterol oxidase catalyzes the oxidation and isomerization of cholesterol to cholest-4-en-3-one. An asparagine residue (Asn485) at the active site is believed to play an important role in catalysis. To test the precise role of Asn485, we mutated it to a leucine and carried out kinetic and crystallographic studies. Steady-state kinetic analysis revealed a 1300-fold decrease in the oxidation k(cat)/K(m) for the mutant enzyme whereas the k(cat)/K(m) for isomerization is only 60-fold slower. The primary kinetic isotope effect in the mutant-catalyzed reaction indicates that 3alpha-H transfer remains the rate-determining step. Measurement of the reduction potentials for the wild-type and N485L enzymes reveals a 76 mV decrease in the reduction potential of the FAD for the mutant enzyme relative to wild type. The crystal structure of the mutant, determined to 1.5 A resolution, reveals a repositioning of the side chain of Met122 near Leu485 to form a hydrophobic pocket. Furthermore, the movement of Met122 facilitates the binding of an additional water molecule, possibly mimicking the position of the equatorial hydroxyl group of the steroid substrate. The wild-type enzyme shows a novel N-H...pi interaction between the side chain of Asn485 and the pyrimidine ring of the cofactor. The loss of this interaction in the N485L mutant destabilizes the reduced flavin and accounts for the decreased reduction potential and rate of oxidation. Thus, the observed structural rearrangement of residues at the active site, as well as the kinetic data and thermodynamic data for the mutant, suggests that Asn485 is important for creating an electrostatic potential around the FAD cofactor enhancing the oxidation reaction.
- Department of Molecular, Sinsheimer Laboratory, University of California, Santa Cruz, Santa Cruz, California 95064, USA.
Organizational Affiliation: