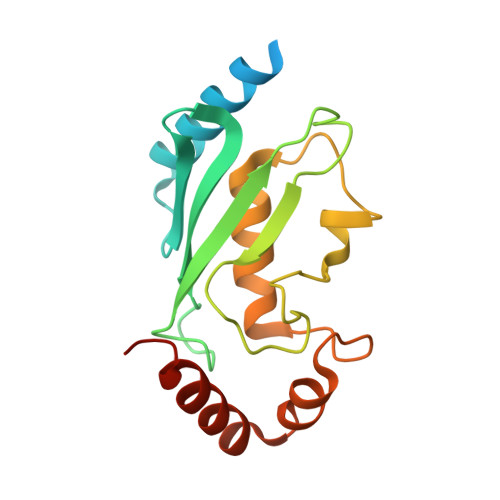
Structural and functional analysis of the human mitotic-specific ubiquitin-conjugating enzyme, UbcH10.
Lin, Y., Hwang, W.C., Basavappa, R.(2002) J Biological Chem 277: 21913-21921
- PubMed: 11927573 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M109398200
- Primary Citation Related Structures:
1I7K - PubMed Abstract:
Cell cycle progression is controlled at several different junctures by the targeted destruction of cell cycle regulatory proteins. These carefully orchestrated events include the destruction of the securin protein to permit entry into anaphase, and the destruction of cyclin B to permit exit from mitosis. These destruction events are mediated by the ubiquitin/proteasome system. The human ubiquitin-conjugating enzyme, UbcH10, is an essential mediator of the mitotic destruction events. We report here the 1.95-A crystal structure of a mutant UbcH10, in which the active site cysteine has been replaced with a serine. Functional analysis indicates that the mutant is active in accepting ubiquitin, although not as efficiently as wild-type. Examination of the crystal structure reveals that the NH2-terminal extension in UbcH10 is disordered and that a conserved 3(10)-helix places a lysine residue near the active site. Analysis of relevant mutants demonstrates that for ubiquitin-adduct formation the presence or absence of the NH2-terminal extension has little effect, whereas the lysine residue near the active site has significant effect. The structure provides additional insight into UbcH10 function including possible sites of interaction with the anaphase promoting complex/cyclosome and the disposition of a putative destruction box motif in the structure.
- Department of Biochemistry and Biophysics, University of Rochester Medical Center, Rochester, New York 14642, USA.
Organizational Affiliation: