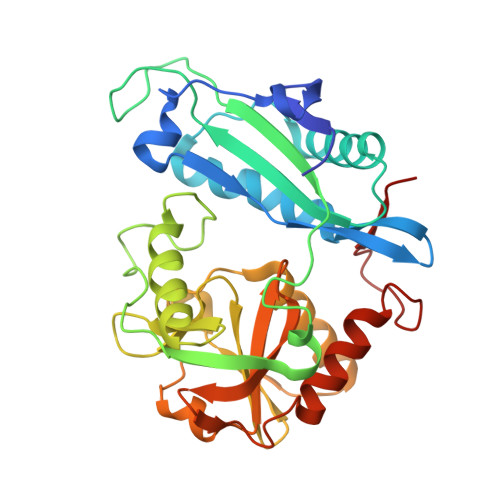
Structures of Escherichia coli branched-chain amino acid aminotransferase and its complexes with 4-methylvalerate and 2-methylleucine: induced fit and substrate recognition of the enzyme.
Okada, K., Hirotsu, K., Hayashi, H., Kagamiyama, H.(2001) Biochemistry 40: 7453-7463
- PubMed: 11412098 Search on PubMed
- DOI: https://doi.org/10.1021/bi010384l
- Primary Citation Related Structures:
1I1K, 1I1L, 1I1M - PubMed Abstract:
The following three-dimensional structures of three forms of Escherichia coli branched-chain amino acid aminotransferase (eBCAT) have been determined by the X-ray diffraction method: the unliganded pyridoxal 5'-phosphate (PLP) form at a 2.1 A resolution, and the two complexes with the substrate analogues, 4-methylvalerate (4-MeVA) as the Michaelis complex model and 2-methylleucine (2-MeLeu) as the external aldimine model at 2.4 A resolution. The enzyme is a trimer of dimers, and each subunit consists of small and large domains, and the interdomain loop. The active site is formed by the residues at the domain interface and those from two loops of the other subunit of the dimer unit, and binds one PLP with its re-face directed toward the protein side. Upon binding of a substrate, Arg40 changes its side-chain direction to interact with the interdomain loop, and the loop, which is disordered in the unliganded form, shows its ordered structure on the active-site cavity, interacts with the hydrophobic side chain of the substrate, and shields it from the solvent region. The substrate binds to the active-site pocket with its alpha-hydrogen toward the protein side, its side-chain on the side of O3 of PLP, and its alpha-carboxylate on the side of the phosphate group of PLP. The hydrophobic side-chain of the substrate is recognized by Phe36, Trp126, Tyr129, Tyr164, Tyr31*, and Val109*. The alpha-carboxylate of the substrate binds to the unique site constructed by three polar groups (two main-chain NH groups of the beta-turn at Thr257 and Ala258 and the hydroxy group of Tyr95) which are activated by the access of Arg40 to the main-chain C=O group of the beta-turn and the coordination of Arg97 to the hydroxy group. Since Arg40 is the only residue that significantly changes its side-chain conformation and directly interacts with the interdomain loop and the beta-turn, the residue plays important roles in the induced fit of the interdomain loop and the alpha-carboxylate recognition of the substrate.
- Department of Chemistry, Graduate School of Science, Osaka City University, Sugimoto, Sumiyoshi-ku, Osaka 558-8585, Japan.
Organizational Affiliation: