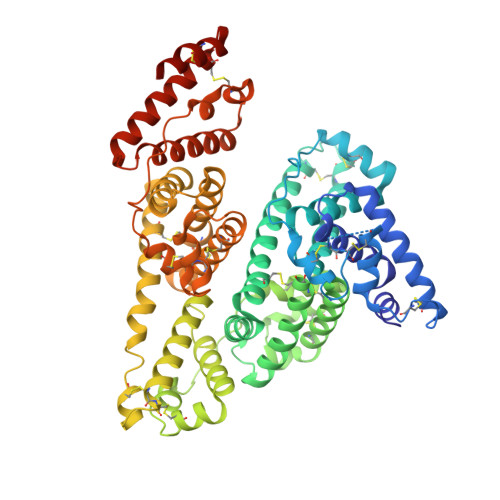
Structural Basis of Albumin-Thyroxine Interactions and Familial Dysalbuminemic Hyperthyroxinemia
Petitpas, I., Petersen, C.E., Ha, C.E., Bhattacharya, A.A., Zunszain, P.A., Ghuman, J., Bhagavan, N.V., Curry, S.(2003) Proc Natl Acad Sci U S A 100: 6440
- PubMed: 12743361
- DOI: https://doi.org/10.1073/pnas.1137188100
- Primary Citation of Related Structures:
1HK1, 1HK2, 1HK3, 1HK4, 1HK5 - PubMed Abstract:
Human serum albumin (HSA) is the major protein component of blood plasma and serves as a transporter for thyroxine and other hydrophobic compounds such as fatty acids and bilirubin. We report here a structural characterization of HSA-thyroxine interactions. Using crystallographic analyses we have identified four binding sites for thyroxine on HSA distributed in subdomains IIA, IIIA, and IIIB. Mutation of residue R218 within subdomain IIA greatly enhances the affinity for thyroxine and causes the elevated serum thyroxine levels associated with familial dysalbuminemic hyperthyroxinemia (FDH). Structural analysis of two FDH mutants of HSA (R218H and R218P) shows that this effect arises because substitution of R218, which contacts the hormone bound in subdomain IIA, produces localized conformational changes to relax steric restrictions on thyroxine binding at this site. We have also found that, although fatty acid binding competes with thyroxine at all four sites, it induces conformational changes that create a fifth hormone-binding site in the cleft between domains I and III, at least 9 A from R218. These structural observations are consistent with binding data showing that HSA retains a high-affinity site for thyroxine in the presence of excess fatty acid that is insensitive to FDH mutations.
- Biophysics Section, Department of Biological Sciences, Imperial College London, South Kensington Campus, United Kingdom.
Organizational Affiliation: