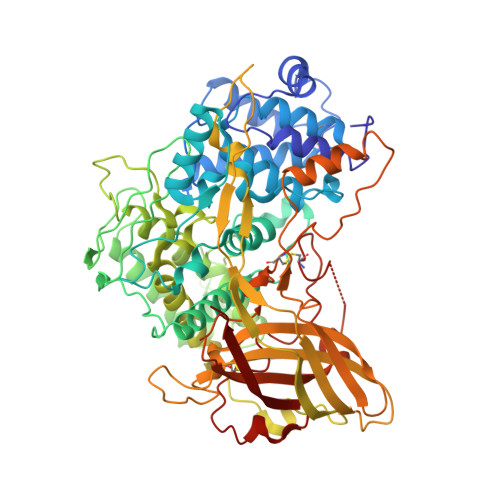
Crystal structure of hexameric haemocyanin from Panulirus interruptus refined at 3.2 A resolution.
Volbeda, A., Hol, W.G.(1989) J Mol Biology 209: 249-279
- PubMed: 2585484
- DOI: https://doi.org/10.1016/0022-2836(89)90276-3
- Primary Citation Related Structures:
1HC1, 1HCY - PubMed Abstract:
The use of non-crystallographic symmetry restraints in the refinement of the haemocyanin hexamer from Panulirus interruptus at 3.2 A resolution has resulted in a final model with a very reasonable geometry and a crystallographic R-factor of 20.1%, using 59,193 observed structure factor amplitudes between 8.0 and 3.2 A. The mean co-ordinate error is approximately 0.35 A. The six subunits appear to be related by symmetry operations that differ slightly from 32 point group symmetry. The six subunits have essentially maintained the same structure. The hexamer, with point group 32, is best described as a trimer of "tight dimers". The contacts between the subunits in such a dimer are more numerous, and better conserved during evolution than contacts in a trimer. The interface of a tight dimer is separated by an internal cavity into two "contact areas". The contact area nearest to the centre of the hexamer is most extensive and consists mainly of residues that are quite conserved among arthropodan haemocyanins. All these residues are provided by the second domain of each subunit. Hence, this second domain may play a crucial role in the allosteric functioning of this oxygen transport protein. The dinuclear copper oxygen-binding site resides in the centre of domain 2. This oxygen-binding centre is not fully accessible from the solvent. Three large cavities occur, however, within each subunit at the interfaces of the three domains. All three cavities contain ordered water molecules, and two of them are accessible from the surrounding solvent. These cavities may play a role in facilitating fast movement of dioxygen towards the binding site, which is situated in a highly conserved, rather hydrophobic core. A detailed definition of the geometry of the copper site is, of course, not possible at the limited resolution of 3.2 A. Nevertheless, it is possible to conclude that each copper is co-ordinated by two, more or less tightly bound, histidine ligands and one more distant histidine residue. The six histidine residues utilize their N epsilon atoms for copper co-ordination, while their N delta atoms are engaged in hydrogen bonds with conserved residues or water molecules. The two distant histidine ligands are located in apical positions and are on opposite sides with respect to the plane approximately defined by the four more tightly bound histidine ligands and the two copper ions. The copper-to-copper distance is 3.5 to 3.6 A in four of the subunits, but this distance deviates considerably in two others.(ABSTRACT TRUNCATED AT 400 WORDS)
- Department of Chemistry, University of Groningen, The Netherlands.
Organizational Affiliation: