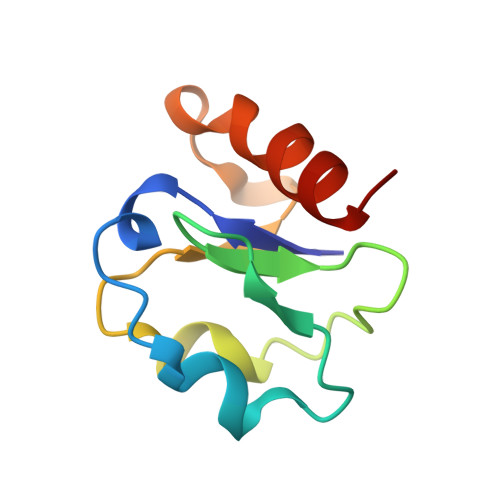
New Insights Into the Thermostability of Bacterial Ferredoxins: High-Resolution Crystal Structure of the Seven-Iron Ferredoxin from Thermus Thermophilus
Macedo-Ribeiro, S., Martins, B.M., Pereira, P.J.B., Buse, G., Huber, R., Soulimane, T.(2001) J Biol Inorg Chem 6: 663
- PubMed: 11681700 Search on PubMed
- DOI: https://doi.org/10.1007/s007750100243
- Primary Citation Related Structures:
1H98 - PubMed Abstract:
The crystal structure of the seven-iron ferredoxin from Thermus thermophilus (FdTt) has been determined at 1.64 A resolution, allowing us to unveil the common mechanisms of thermostabilization within "bacterial-type" ferredoxins. FdTt and other homologous thermophilic seven-iron ferredoxins are smaller than their mesophilic counterparts. Thermostabilizing features are optimized in a minimal structural and functional unit, with an extensive cross-linking of secondary structure elements mediated by improved polar and hydrophobic interactions. Most of the potentially stabilizing features are focused on the vicinity of the functional [3Fe-4S] cluster. The structural [4Fe-4S] cluster is shielded in thermophilic FdTt by an increased number of polar interactions involving the two N-terminal residues. Comparisons with the hyperthermostable ferredoxin from Thermotoga maritima reveal that (1) a reduction in the number of non-glycine residues in strained conformations, (2) improved polar interactions within the common iron-sulfur cluster binding (betaalphabeta)2 motif, and (3) an optimized charge distribution at the protein surface, constitute a common strategy for increasing the thermal stability of these ferredoxins.
- Max-Planck-Institut für Biochemie, Abteilung Strukturforschungm, Martinsried, Germany. sandra@aramis.ibmc.up.pt
Organizational Affiliation: