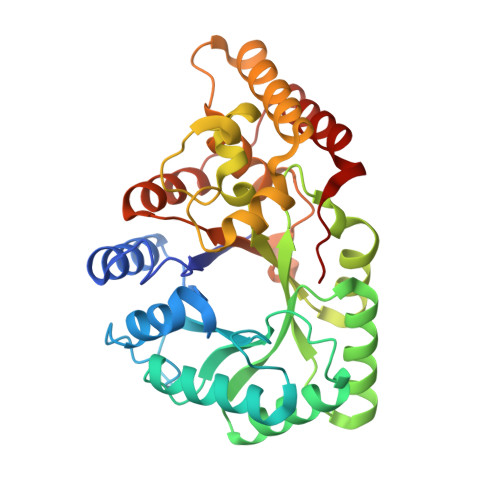
The Crystal Structure of Rat Liver Akr7A1: A Dimeric Member of the Aldo-Keto Reductase Superfamily
Kozma, E., Brown, E., Ellis, E.M., Lapthorn, A.J.(2002) J Biological Chem 277: 16285
- PubMed: 11839745 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M110808200
- Primary Citation Related Structures:
1GVE - PubMed Abstract:
The structure of the rat liver aflatoxin dialdehyde reductase (AKR7A1) has been solved to 1.38-A resolution. Although it shares a similar alpha/beta-barrel structure with other members of the aldo-keto reductase superfamily, AKR7A1 is the first dimeric member to be crystallized. The crystal structure also reveals details of the ternary complex as one subunit of the dimer contains NADP(+) and the inhibitor citrate. Although the underlying catalytic mechanism appears similar to other aldo-keto reductases, the substrate-binding pocket contains several charged amino acids (Arg-231 and Arg-327) that distinguish it from previously characterized aldo-keto reductases with respect to size and charge. These differences account for the substrate specificity for 4-carbon acid-aldehydes such as succinic semialdehyde and 2-carboxybenzaldehyde as well as for the idiosyncratic substrate aflatoxin B(1) dialdehyde of this subfamily of enzymes. Structural differences between the AKR7A1 ternary complex and apoenzyme reveal a significant hinged movement of the enzyme involving not only the loops of the structure but also parts of the alpha/beta-barrel most intimately involved in cofactor binding.
- Department of Chemistry, University of Glasgow, Glasgow G12 8QQ, United Kingdom.
Organizational Affiliation: