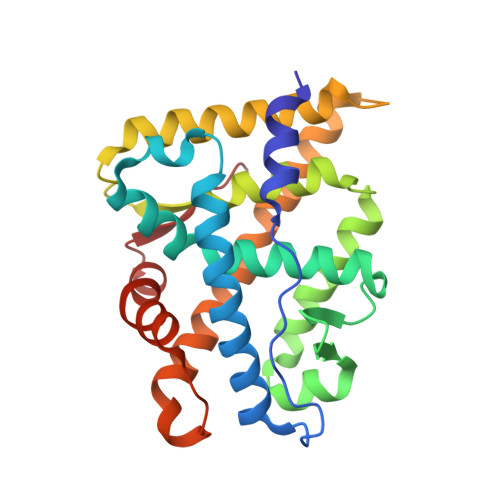
Structural Basis for the Glucocorticoid Response in a Mutant Human Androgen Receptor (Ar(Ccr)) Derived from an Androgen-Independent Prostate Cancer
Matias, P.M., Carrondo, M.A., Coelho, R., Thomaz, M., Zhao, X.-Y., Wegg, A., Crusius, K., Egner, U., Donner, P.(2002) J Med Chem 45: 1439-1446
- PubMed: 11906285 Search on PubMed
- DOI: https://doi.org/10.1021/jm011072j
- Primary Citation Related Structures:
1GS4 - PubMed Abstract:
The crystal structure of a mutant androgen receptor (AR) ligand-binding domain (LBD) in complex with the agonist 9alpha-fluorocortisol has been determined at 1.95 A resolution. This mutant AR contains two mutations (L701H and T877A) and was previously reported as a high-affinity cortisol/cortisone responsive AR (AR(ccr)) isolated from the androgen-independent human prostate cancer cell lines MDA PCa 2a and 2b (Zhao et al. Nature Med. 2000, 6, 703-6). The three-dimensional structure of the AR(ccr) LBD complexed with 9alpha-fluorocortisol shows the typical conformation of an agonist-bound nuclear receptor in which helix 12 is precisely positioned as a "lid" for the ligand-binding pocket. Binding of 9alpha-fluorocortisol to the AR(ccr) involves favorable hydrogen bond patterns on the C17 and C21 substituents of the ligand due to the mutations at 701 and 877 in the AR(ccr). Our studies provide the first structural explanation for the glucocorticoid activation of AR(ccr), which is important for the development of new therapeutic treatments for androgen-independent prostate cancer.
- Instituto de Tecnologia Química e Biológica, Universidade Nova de Lisboa, Apartado 127, 2780 Oeiras, Portugal.
Organizational Affiliation: