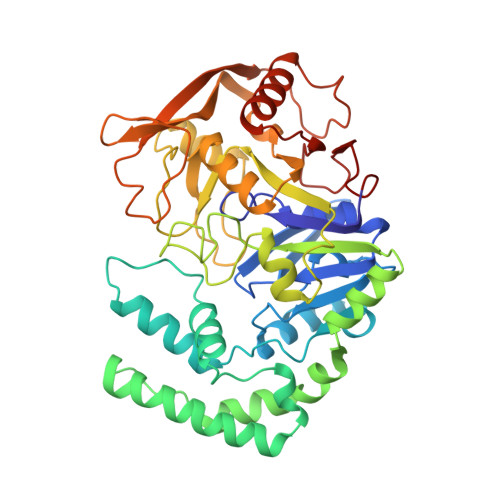
Crystal structures of adenylosuccinate synthetase from Escherichia coli complexed with GDP, IMP hadacidin, NO3-, and Mg2+.
Poland, B.W., Fromm, H.J., Honzatko, R.B.(1996) J Mol Biology 264: 1013-1027
- PubMed: 9000627
- DOI: https://doi.org/10.1006/jmbi.1996.0693
- Primary Citation of Related Structures:
1GIM, 1GIN - PubMed Abstract:
Crystal structures of adenylosuccinate synthetase from Esherichia coli complexed with Mg2+, IMP, GDP, NO3- and hadacidin at 298 and 100 K have been refined to R-factors of 0.188 and 0.206 against data to 2.8 A and 2.5 A resolution, respectively. Conformational changes of up to 9 A relative to the unligated enzyme occur in loops that bind to Mg2+, GDP, IMP and hadacidin. Mg2+ binds directly to GDP, NO3-, hadacidin and the protein, but is only five-coordinated. Asp13, which approaches, but does not occupy the sixth coordination site of Mg2+, hydrogen bonds to N1 of IMP. The nitrogen atom of NO3- is approximately 2.7 A from O6 of IMP, reflecting a strong electrostatic interaction between the electron-deficient nitrogen atom and the electron-rich O6. The spatial relationships between GDP, NO3- and Mg2+ suggest an interaction between the beta,gamma-bridging oxygen atom of GTP and Mg2+ in the enzyme-substrate complex. His41 hydrogen bonds to the beta-phosphate group of GDP and approaches bound NO3-. The aldehyde group of hadacidin coordinates to the Mg2+, while its carboxyl group interacts with backbone amide groups 299 to 303 and the side-chain of Arg303. The 5'-phosphate group of IMP interacts with Asn38, Thr129, Thr239 and Arg143 (from a monomer related by 2-fold symmetry). A mechanism is proposed for the two-step reaction governed by the synthetase, in which His41 and Asp13 are essential catalytic side-chains.
- Department of Biochemistry and Biophysics, Iowa State University, Ames 50011, USA.
Organizational Affiliation: