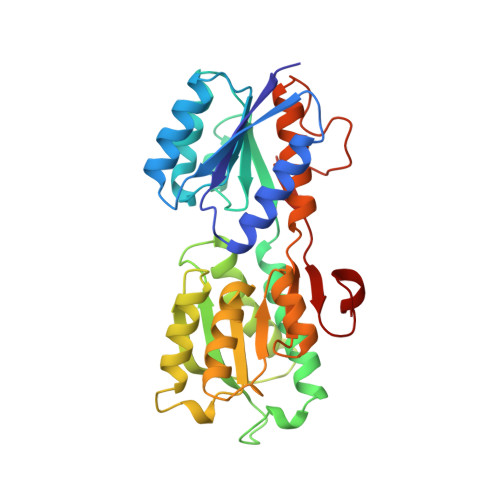
The 1.7 A refined X-ray structure of the periplasmic glucose/galactose receptor from Salmonella typhimurium.
Zou, J.Y., Flocco, M.M., Mowbray, S.L.(1993) J Mol Biology 233: 739-752
- PubMed: 8240551 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1993.1549
- Primary Citation Related Structures:
1GCA - PubMed Abstract:
The X-ray structure of the periplasmic glucose/galactose receptor (binding protein) of Salmonella typhimurium (GBP-S) has been refined at 1.7 A resolution with an R-factor of 19.0%. The model contains all 309 residues of the amino acid sequence, 153 water molecules, a calcium ion and beta-D-galactose. The protein consists of two very similar structural domains, each of which is composed a core of parallel beta-sheet flanked on both sides by alpha-helices. Three short stretches of amino acid chain (from symmetrically related portions of the structure) link the domains, and presumably act as a hinge to allow their relative movement in functionally important conformational changes. Galactose is bound between the domains, interacting with a number of side-chains from the loops lining the binding cleft. A combination of hydrogen bonding, hydrophobic and steric effects give rise to tight binding (dissociation constant 0.2 microM) and high specificity. Of nine hydrogen bonding groups, three are aspartate, three asparagine, one histidine (unprotonated), one arginine and one water, contributing 13 hydrogen bonds in total. Additional residues pack against (primarily) non-polar faces of the sugar molecule. The precise arrangement of the hydrogen bonding and hydrophobic residues results in an enclosed binding site with a shape that is a composite of those of the allowed sugar molecules. It is presumed that ligands bind to a more open form of the receptor that then closes by rotation in the hinge. Comparison with the GBP-S structure solved earlier in complex with glucose shows no significant changes, even for the aspartate residue most directly involved with the different sugars. Comparison with the galactose/glucose receptor of Escherichia coli indicates that these two proteins are very similar in overall structure, with the main difference being a 2 to 3 degrees rotation in the hinge. This difference appears to be the result of different crystal packing for the two proteins; it is likely that both conformations are normally found in solution.
- Department of Molecular Biology, Uppsala University, Sweden.
Organizational Affiliation: