Kinetic and structural characterization of mutations of glycine 216 in alpha-lytic protease: a new target for engineering substrate specificity.
Mace, J.E., Agard, D.A.(1995) J Mol Biology 254: 720-736
- PubMed: 7500345 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1995.0650
- Primary Citation Related Structures:
1GBA, 1GBB, 1GBC, 1GBD, 1GBE, 1GBF, 1GBH, 1GBI, 1GBJ, 1GBK, 1GBL, 1GBM - PubMed Abstract:
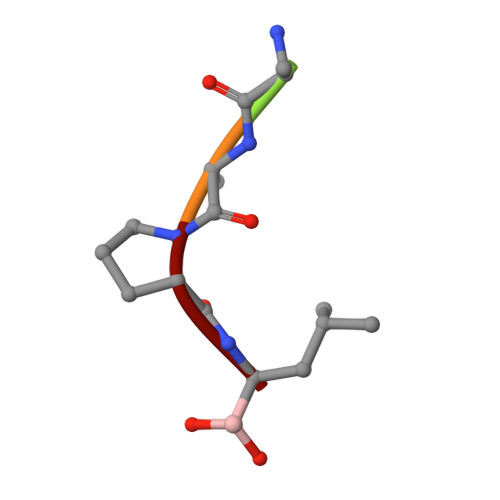
Gly216 in the active site of the broadly specific MA190 mutant of alpha-lytic protease has been found to be remarkably tolerant of amino acid substitutions. Side-chains as large as Trp can be accommodated within the substrate-binding pocket without abolishing catalysis, and have major effects upon the substrate specificity of the enzyme. Kinetic characterization of eleven enzymatically active mutants against a panel of eight substrates clearly revealed the functional consequences of the substitutions at position 216. To understand better the structural basis for their altered specificity, the GA216 + MA190 and GL216 + MA190 mutants have been crystallized both with and without a representative series of peptide boronic acid transition-state analog inhibitors. An empirical description and non-parametric statistical analysis of structural variation among these enzyme: inhibitor complexes is presented. The roles of active site plasticity and dynamics in alpha-lytic protease function and substrate preference are also addressed. The results strongly suggest that substrate specificity determination in alpha-lytic protease is a distributed property of the active site and substrate molecule.
- Howard Hughes Medical Institute, University of California, San Francisco 94143-0448, USA.
Organizational Affiliation: