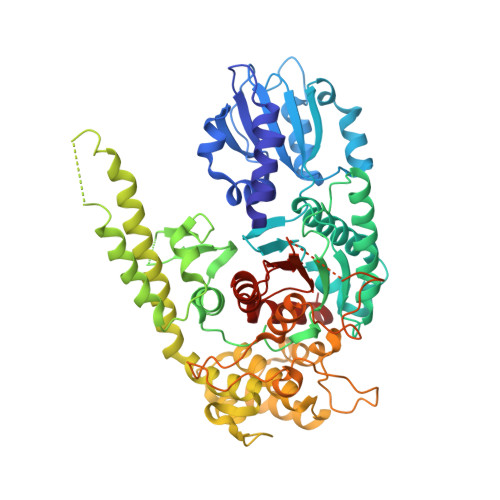
Crystal structures of neuronal squid Sec1 implicate inter-domain hinge movement in the release of t-SNAREs.
Bracher, A., Weissenhorn, W.(2001) J Mol Biology 306: 7-13
- PubMed: 11178889
- DOI: https://doi.org/10.1006/jmbi.2000.4347
- Primary Citation of Related Structures:
1FVF, 1FVH - PubMed Abstract:
Sec1 molecules associate with t-SNAREs from the syntaxin family in a heterodimeric complex that plays an essential role in vesicle transport and membrane fusion. Neuronal rat n-Sec1 has an arch-shaped three-domain structure, which binds syntaxin 1a through contacts in domains 1 and 3. In both rat nSec1 and homologous squid s-Sec1, a potential effector-molecule binding-pocket is shaped by residues from domains 1 and 2 and is localized on the opposite side of the syntaxin 1a interaction site. Comparison of several crystal forms of unliganded neuronal squid Sec1 indicates a hinge region between domains 1 and 2 which allows domain 1 to rotate along a central axis. This movement could release syntaxin 1a upon interaction with a yet unspecified Sec1 effector molecule(s). The binding of an effector protein may also directly affect the conformation of the helical hairpin of domain 3, which contributes the other significant syntaxin 1a binding sites in the rat nSec1/syntaxin 1a complex structure but adopts multiple conformations in the unliganded s-Sec1 structures reported here.
- European Molecular Biology Laboratory (EMBL), 6 rue Jules Horowitz, Grenoble, 38000, France.
Organizational Affiliation: