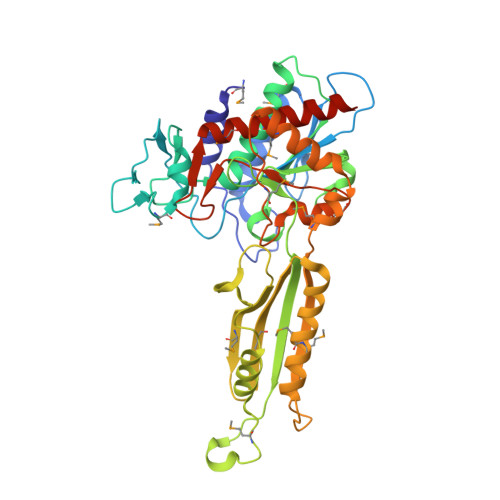
Structure of Peptidase T from Salmonella typhimurium
Hakansson, K., Miller, C.G.(2002) Eur J Biochem 269: 443-450
- PubMed: 11856302
- DOI: https://doi.org/10.1046/j.0014-2956.2001.02665.x
- Primary Citation Related Structures:
1FNO - PubMed Abstract:
The structure of peptidase T, or tripeptidase, was determined by multiple wavelength anomalous dispersion (MAD) methodology and refined to 2.4 A resolution. Peptidase T comprises two domains; a catalytic domain with an active site containing two metal ions, and a smaller domain formed through a long insertion into the catalytic domain. The two metal ions, presumably zinc, are separated by 3.3 A, and are coordinated by five carboxylate and histidine ligands. The molecular surface of the active site is negatively charged. Peptidase T has the same basic fold as carboxypeptidase G2. When the structures of the two enzymes are superimposed, a number of homologous residues, not evident from the sequence alone, could be identified. Comparison of the active sites of peptidase T, carboxypeptidase G2, Aeromonas proteolytica aminopeptidase, carboxypeptidase A and leucine aminopeptidase reveals a common structural framework with interesting similarities and differences in the active sites and in the zinc coordination. A putative binding site for the C-terminal end of the tripeptide substrate was found at a peptidase T specific fingerprint sequence motif.
- Department of Microbiology, University of Illinois at Urbana-Champaign, Urbana, IL 61801, USA.
Organizational Affiliation: