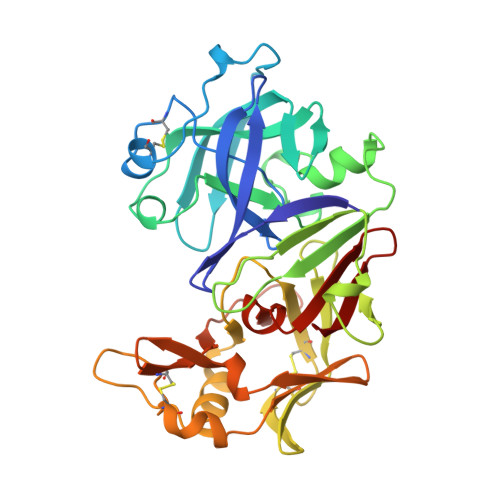
Structure of human uropepsin at 2.45 A resolution.
Canduri, F., Teodoro, L.G., Fadel, V., Lorenzi, C.C., Hial, V., Gomes, R.A., Neto, J.R., de Azevedo, W.F.(2001) Acta Crystallogr D Biol Crystallogr 57: 1560-1570
- PubMed: 11679720 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444901013865
- Primary Citation Related Structures:
1FLH - PubMed Abstract:
The molecular structure of human uropepsin, an aspartic proteinase from the urine produced in the form of pepsinogen A in the gastric mucosa, has been determined by molecular replacement using human pepsin as the search model. Crystals belong to space group P2(1)2(1)2(1), with unit-cell parameters a = 50.99, b = 75.56, c = 89.90 A. Crystallographic refinement led to an R factor of 0.161 at 2.45 A resolution. The positions of 2437 non-H protein atoms in 326 residues have been determined and the model contains 143 water molecules. The structure is bilobal, consisting of two predominantly beta-sheet lobes which, as observed in other aspartic proteinases, are related by a pseudo-twofold axis. A model of the uropepsin-pepstatin complex has been constructed based on the high-resolution crystal structure of pepsin complexed with pepstatin.
- Departamento de Física, IBILCE, UNESP, São José do Rio Preto, SP 15054-000, Brazil.
Organizational Affiliation: