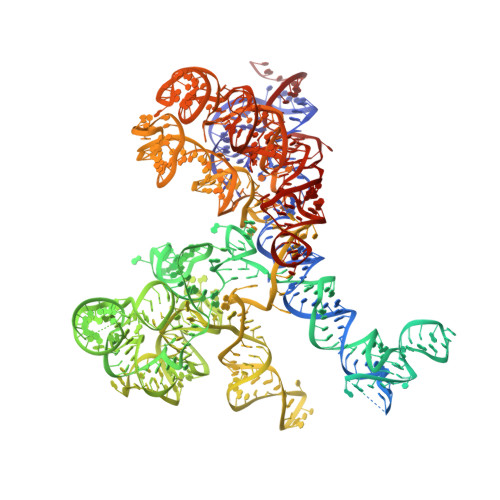
The structural basis of ribosome activity in peptide bond synthesis.
Nissen, P., Hansen, J., Ban, N., Moore, P.B., Steitz, T.A.(2000) Science 289: 920-930
- PubMed: 10937990 Search on PubMed
- DOI: https://doi.org/10.1126/science.289.5481.920
- Primary Citation Related Structures:
1FFZ, 1FG0 - PubMed Abstract:
Using the atomic structures of the large ribosomal subunit from Haloarcula marismortui and its complexes with two substrate analogs, we establish that the ribosome is a ribozyme and address the catalytic properties of its all-RNA active site. Both substrate analogs are contacted exclusively by conserved ribosomal RNA (rRNA) residues from domain V of 23S rRNA; there are no protein side-chain atoms closer than about 18 angstroms to the peptide bond being synthesized. The mechanism of peptide bond synthesis appears to resemble the reverse of the acylation step in serine proteases, with the base of A2486 (A2451 in Escherichia coli) playing the same general base role as histidine-57 in chymotrypsin. The unusual pK(a) (where K(a) is the acid dissociation constant) required for A2486 to perform this function may derive in part from its hydrogen bonding to G2482 (G2447 in E. coli), which also interacts with a buried phosphate that could stabilize unusual tautomers of these two bases. The polypeptide exit tunnel is largely formed by RNA but has significant contributions from proteins L4, L22, and L39e, and its exit is encircled by proteins L19, L22, L23, L24, L29, and L31e.
- Department of Molecular Biophysics and Biochemistry and Department of Chemistry, Yale University, and Howard Hughes Medical Institute, New Haven, CT 06520-8114, USA.
Organizational Affiliation: