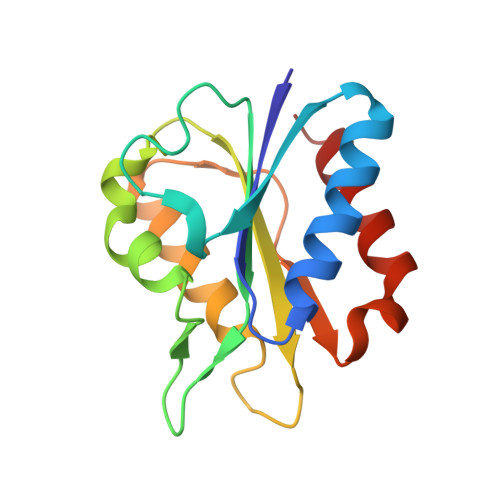
Structures and comparison of the Y98H (2.0 A) and Y98W (1.5 A) mutants of flavodoxin (Desulfovibrio vulgaris).
Reynolds, R.A., Watt, W., Watenpaugh, K.D.(2001) Acta Crystallogr D Biol Crystallogr 57: 527-535
- PubMed: 11264581
- DOI: https://doi.org/10.1107/s0907444901002554
- Primary Citation Related Structures:
1F4P, 1I1O - PubMed Abstract:
The structures for two mutants at the Tyr98 site of Desulfovibrio vulgaris flavodoxin have been determined. The first, a tyrosine-to-histidine (Y98H) variant, was determined at the moderately high resolution of 2.0 A, while the tyrosine-to-tryptophan variant (Y98W) yielded very high resolution data (beyond 1.5 A) allowing a detailed look at the water structure, alternate side-chain conformations and the planarity of the FMN. Both structures were solved by molecular replacement beginning with the native (P2A) coordinates as a starting point. The Y98H variant of D. vulgaris flavodoxin crystallizes in space group P2(1)2(1)2(1), with unit-cell parameters a = 41.96, b = 61.45, c = 57.04 A, while the Y98W mutant adopts space group P2(1), with a = 41.29, b = 55.82, c = 32.52 A, beta = 100.68 degrees. Refinement for both mutants utilized PROLSQ followed by, for the high-resolution Y98W structure, anisotropic refinement as implemented in SHELXL. Final R factors of 17% for the Y98H mutant and 9.8% for the Y98W mutant were obtained. For the high-resolution (1.5 A) Y98W mutant, 31,010 unique reflections were collected from a single crystal. The final model includes 273 solvent molecules, with eight side chains assuming multiple conformations. At this resolution, the detailed conformation of the FMN can be observed, with both a bow and twist being noted. A comparison is made between the two mutants and the different oxidation states of the native flavodoxin. Although both mutants show similar E(2) (oxidized/semiquinone) one-electron redox potentials to the native, the E(1) (semiquinone/hydroquinone) redox potential for the Y98H mutant is significantly different from that of the Y98W variant and the native protein. The surprising similarity in the folding of the polypeptide chain 60--64 between the two mutants and the reduced states of the native is discussed. The interaction between O61 and N5 in the flavin is discussed because of the new conformation of this loop.
- Physics Department, Grand Valley State University, Allendale, Michigan, USA.
Organizational Affiliation: