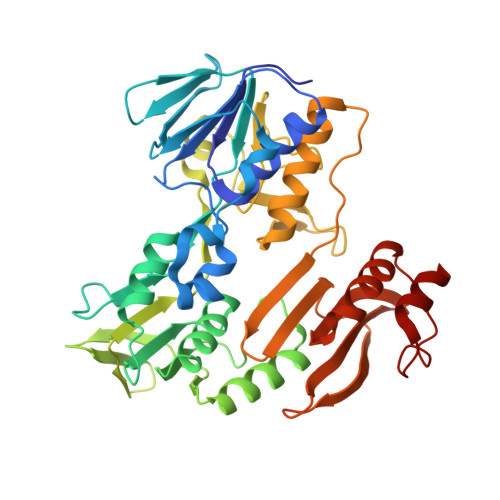
Crystal structure of NADH-dependent ferredoxin reductase component in biphenyl dioxygenase.
Senda, T., Yamada, T., Sakurai, N., Kubota, M., Nishizaki, T., Masai, E., Fukuda, M., Mitsuidagger, Y.(2000) J Mol Biology 304: 397-410
- PubMed: 11090282
- DOI: https://doi.org/10.1006/jmbi.2000.4200
- Primary Citation Related Structures:
1D7Y, 1F3P - PubMed Abstract:
Oxidative biodegradation of aromatic compounds by bacteria usually begins with hydroxylation of the aromatic ring by multi-component dioxygenases like benzene dioxygenase, biphenyl dioxygenase, and others. These enzymes are composed of ferredoxin reductase, ferredoxin, and terminal oxygenase. Reducing equivalents that originate from NADH are transferred from ferredoxin reductase to ferredoxin and, in turn, to the terminal oxygenase, thus resulting in the activation of a dioxygen. BphA4 is the ferredoxin reductase component of biphenyl dioxygenase from Pseudomonas sp. strain KKS102. The amino acid sequence of BphA4 exhibits significant homology with the putidaredoxin reductase of the cytochrome P450cam system in Pseudomonas putida, as well as with various other oxygenase-coupled NADH-dependent ferredoxin reductases (ONFRs) of bacteria. To date, no structural information has been provided for the ferredoxin reductase component of the dioxygenase systems. In order to provide a structural basis for discussing the mechanism of electron transport between ferredoxin reductase and ferredoxin, crystal structures of BphA4 and its NADH complex were solved. The three-dimensional structure of BphA4 is different from those of ferredoxin reductases whose structures have already been determined, but adopts essentially the same fold as the enzymes of the glutathione reductase (GR) family. Also the three-dimensional structure of the first two domains of BphA4 adopts a fold similar to that of adrenodoxin reductase (AdR) in the mitochondrial cytochrome P450 system. Comparing the amino acid sequence with what is known of the three-dimensional structure of BphA4 strongly suggests that the other ONFRs have secondary structural features that are similar to that of BphA4. This analysis of the crystal structures of BphA4 suggests that Lys53 and Glu159 seem to be involved in the hydride transfer from NADH to FAD. Since the amino acid residues around the active site, some of which seem to be important to electron transport, are highly conserved among ONFRs, it is likely that the mechanism of electron transport of BphA4 is quite applicable to other ONFRs.
- Division of Protein Engineering, University of Technology, Nagaoka, Niigata, Japan. senda@vos.nagaskaut.ac.jp
Organizational Affiliation: