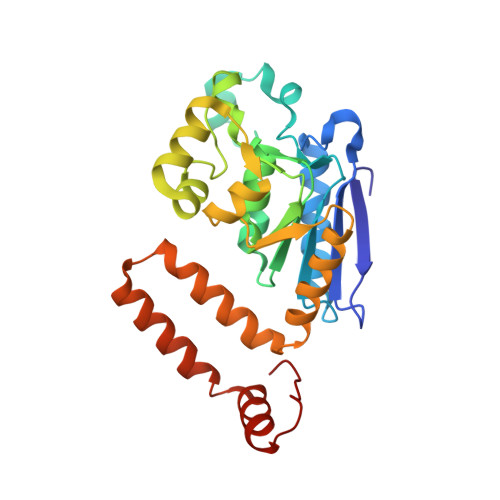
Structural mechanism of enoyl-CoA hydratase: three atoms from a single water are added in either an E1cb stepwise or concerted fashion.
Bahnson, B.J., Anderson, V.E., Petsko, G.A.(2002) Biochemistry 41: 2621-2629
- PubMed: 11851409 Search on PubMed
- DOI: https://doi.org/10.1021/bi015844p
- Primary Citation Related Structures:
1EY3 - PubMed Abstract:
We have determined the crystal structure of the enzyme enoyl-CoA hydratase (ECH) from rat liver with the bound substrate 4-(N,N-dimethylamino)cinnamoyl-CoA using X-ray diffraction data to a resolution of 2.3 A. In addition to the thiolester substrate, the catalytic water, which is added in the hydration reaction, has been modeled into well-defined electron density in each of the six active sites of the physiological hexamer within the crystallographic asymmetric unit. The catalytic water bridges Glu(144) and Glu(164) of the enzyme and has a lone pair of electrons poised to react with C(3) of the enzyme-bound alpha,beta-unsaturated thiolester. The water molecule, which bridges two glutamate residues, is reminiscent of the enolase active site. However, unlike enolase, which has a lysine available to donate a proton, there are no other sources of protons available from other active site residues in ECH. Furthermore, an analysis of the hydrogen-bonding network of the active site suggests that both Glu(144) and Glu(164) are ionized and carry a negative charge with no reasonable place to have a protonated carboxylate. This lack of hydrogen-bonding acceptors that could accommodate a source of a proton, other than from the water molecule, leads to a hypothesis that the three atoms from a single water molecule are added across the double bond to form the hydrated product. The structural results are discussed in connection with details of the mechanism, which have been elucidated from kinetics, site-directed mutagenesis, and spectroscopy of enzyme-substrate species, in presenting an atomic-resolution mechanism of the reaction. Contrary to the previous interpretation, the structure of the E-S complex together with previously determined kinetic isotope effects is consistent with either a concerted mechanism or an E1cb stepwise mechanism.
- Department of Chemistry and Biochemistry, University of Delaware, Newark, Delaware 19716, USA. bahnson@udel.edu
Organizational Affiliation: