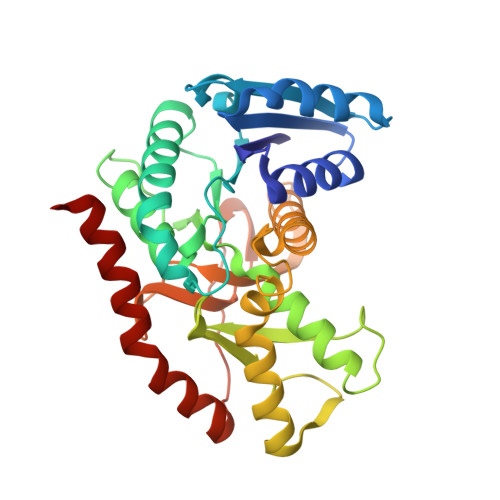
Crystal structure of a ternary complex of Escherichia coli malate dehydrogenase citrate and NAD at 1.9 A resolution.
Hall, M.D., Banaszak, L.J.(1993) J Mol Biology 232: 213-222
- PubMed: 8331658 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1993.1377
- Primary Citation Related Structures:
1EMD - PubMed Abstract:
The structure of malate dehydrogenase from Escherichia coli complexed with the substrate analog, citrate and the cofactor NAD, has been determined by X-ray crystallography. A monoclinic crystal of the malate dehydrogenase, grown in citrate buffer, was soaked in 10 mM NAD solution and found to be isomorphous with the apo-form. The X-ray data extended to 1.9 A, nearly the same resolution limit as the apo-enzyme crystals. The ternary complex of malate dehydrogenase has very few conformational differences from that of the pseudo binary complex of enzyme with bound citrate. In addition, the NAD molecule has a very similar conformation to the NAD as found in the crystal structure of the cytosolic eukaryotic malate dehydrogenase. Similar hydrogen bond interactions are made by both enzymes from polar groups belonging to the NAD. Such interactions include hydrogen bonds from the ribose oxygens and the phosphate oxygens, to backbone amide and carbonyl atoms of the protein and to side-chains of a select few conserved hydrophilic residues. The only notable difference occurs in the active site region where the nicotinamide moiety is obstructed from further entering the active site by the C-6 carbonyl atoms of citrate. In this position there are no direct polar interactions between the protein and the nicotinamide moiety. Energy minimization of the structure with malate substituted for citrate in the active site shows that the nicotinamide moiety assumes the same position in the active site as the NAD in cytosolic malate dehydrogenase. The carboxamide atoms of the energy minimized model make significant hydrogen bond interactions with the catalytic residue, H177, and with the main-chain atoms of I117 and V146 in the vicinity of the active site, while the position of the rest of the cofactor remains unchanged.
- Department of Biochemistry, University of Minnesota, Minneapolis 55455.
Organizational Affiliation: