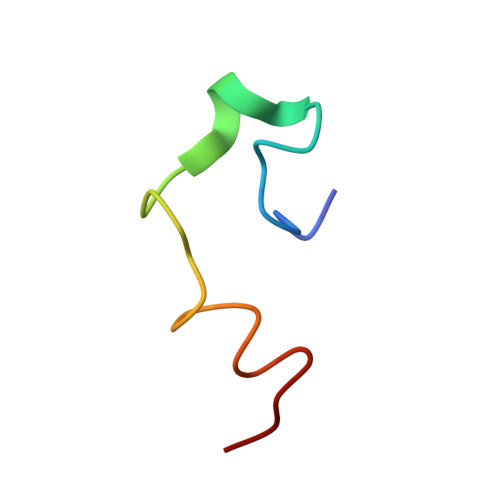
Structures of the intradiskal loops and amino terminus of the G-protein receptor, rhodopsin.
Yeagle, P.L., Salloum, A., Chopra, A., Bhawsar, N., Ali, L., Kuzmanovski, G., Alderfer, J.L., Albert, A.D.(2000) J Pept Res 55: 455-465
- PubMed: 10888202 Search on PubMed
- DOI: https://doi.org/10.1034/j.1399-3011.2000.00707.x
- Primary Citation Related Structures:
1EDS, 1EDV, 1EDW, 1EDX - PubMed Abstract:
The intradiskal surface of the transmembrane protein, rhodopsin, consists of the amino terminal domain and three loops connecting six of the seven transmembrane helices. This surface corresponds to the extracellular surface of other G-protein receptors. Peptides that represent each of the extramembraneous domains on this surface (three loops and the amino terminus) were synthesized. These peptides also included residues which, based on a hydrophobic plot, could be expected to be part of the transmembrane helix. The structure of each of these peptides in solution was then determined using two-dimensional 1H nuclear magnetic resonance. All peptide domains showed ordered structures in solution. The structures of each of the peptides from intradiskal loops of rhodopsin exhibited a turn in the central region of the peptide. The ends of the peptides show an unwinding of the transmembrane helices to form this turn. The amino terminal domain peptide exhibited alpha-helical regions with breaks and bends at proline residues. This region forms a compact domain. Together, the structures for the loop and amino terminus domains indicate that the intradiskal surface of rhodopsin is ordered. These data further suggest a structural motif for short loops in transmembrane proteins. The ordered structures of these loops, in the absence of the transmembrane helices, indicate that the primary sequences of these loops are sufficient to code for the turn.
- Department of Molecular and Cell Biology, University of Connecticut, Storrs 06269, USA. yeagle@uconnvm.uconn.edu
Organizational Affiliation: