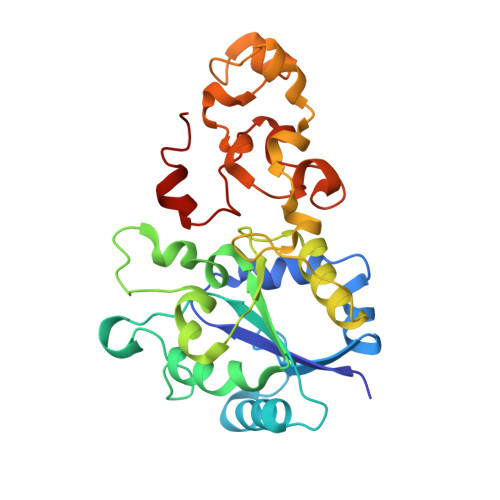
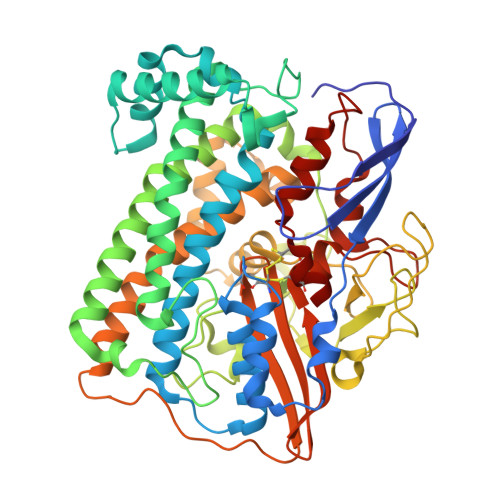
[Nife] Hydrogenase from Desulfovibrio Desulfuricans Atcc 27774: Gene Sequencing, Three-Dimensional Structure Determination and Refinement at 1.8 A and Modelling Studies of its Interaction with the Tetrahaem Cytochrome C3.
Matias, P.M., Soares, C.M., Saraiva, L.M., Coelho, R., Morais, J., Le Gall, J., Carrondo, M.A.(2001) J Biol Inorg Chem 6: 63-81
- PubMed: 11191224
- DOI: https://doi.org/10.1007/s007750000167
- Primary Citation of Related Structures:
1E3D - PubMed Abstract:
The primary and three-dimensional structures of a [NiFe] hydrogenase isolated from D. desulfitricans ATCC 27774 were determined, by nucleotide analysis and single-crystal X-ray crystallography. The three-dimensional structural model was refined to R=0.167 and Rfree=0.223 using data to 1.8 A resolution. Two unique structural features are observed: the [4Fe-4S] cluster nearest the [NiFe] centre has been modified [4Fe-3S-3O] by loss of one sulfur atom and inclusion of three oxygen atoms; a three-fold disorder was observed for Cys536 which binds to the nickel atom in the [NiFe] centre. Also, the bridging sulfur atom that caps the active site was found to have partial occupancy, thus corresponding to a partly activated enzyme. These structural features may have biological relevance. In particular, the two less-populated rotamers of Cys536 may be involved in the activation process of the enzyme, as well as in the catalytic cycle. Molecular modelling studies were carried out on the interaction between this [NiFe] hydrogenase and its physiological partner, the tetrahaem cytochrome c3 from the same organism. The lowest energy docking solutions were found to correspond to an interaction between the haem IV region in tetrahaem cytochrome c3 with the distal [4Fe-4S] cluster in [NiFe] hydrogenase. This interaction should correspond to efficient electron transfer and be physiologically relevant, given the proximity of the two redox centres and the fact that electron transfer decay coupling calculations show high coupling values and a short electron transfer pathway. On the other hand, other docking solutions have been found that, despite showing low electron transfer efficiency, may give clues on possible proton transfer mechanisms between the two molecules.
- Instituto de Tecnologia Química e Biológica, Universidade Nova de Lisboa, Oeiras, Portugal.
Organizational Affiliation: