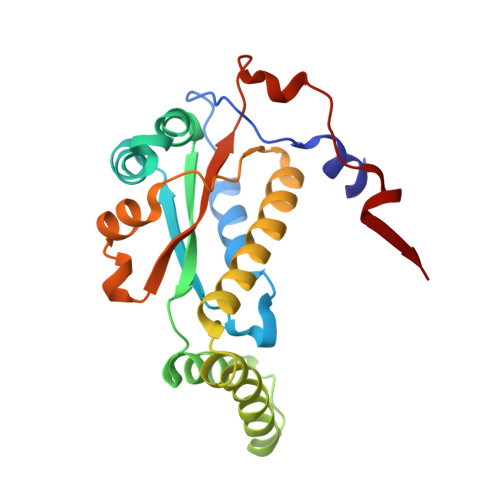
Crystal structure of FMN-dependent nitroreductase from Escherichia coli B: a prodrug-activating enzyme.
Parkinson, G.N., Skelly, J.V., Neidle, S.(2000) J Med Chem 43: 3624-3631
- PubMed: 11020276 Search on PubMed
- DOI: https://doi.org/10.1021/jm000159m
- Primary Citation Related Structures:
1DS7 - PubMed Abstract:
The FMN-dependent flavoprotein nitroreductase from Escherichia coli B (NTR) is used in cancer chemotherapy to activate a range of prodrugs. The crystal structure of this enzyme has been determined, using molecular replacement methods and refined at 2.06 A resolution. The recombinant 24-kDa enzyme was crystallized in the tetragonal space group P4(1)2(1)2, with unit cell dimensions of a = b = 57.74 A and c = 275.51 A and two molecules in the asymmetric unit. The structure has a final R factor of 20.3% (R(free) = 26.7%), for all data between the resolution ranges of 10-2.06 A, and includes 4453 protein atoms, 230 water molecules, and 2 flavin mononucleotide (FMN) molecules. The functional unit is a homodimer, which forms the asymmetric unit in the crystal structure. The tertiary structures of these two monomers and their subunit interactions are nearly identical. The molecular replacement search model, the crystal structure of the major NAD(P)H:FMN oxidoreductase of Vibrio fisheri (FRase 1), was selected on the basis of its high sequence identity to that of NTR. The final superposition of these two enzymes revealed a very similar overall fold, with variation in the structures focused around surface loops and helices near the FMN cofactor. Helix G is implicated in substrate specificity and is better resolved in the present NTR structure than in the previously reported FRase 1 structure. The FMN binding pocket is also well-resolved, showing the presence of two channels leading into the active site. The amino acid side chains and main chain atoms interacting with the FMN are well-ordered. The structure of the substrate binding pocket has been used to examine substrate specificity and enzyme kinetics for prodrugs used in antibody-directed enzyme prodrug therapy (ADEPT) and gene-directed enzyme prodrug therapy (GDEPT).
- CRC Biomolecular Structure Unit, Chester Beatty Laboratories, The Institute of Cancer Research, London SW3 6JB, United Kingdom.
Organizational Affiliation: