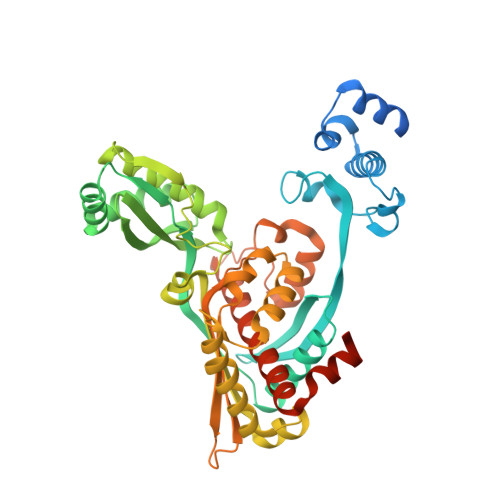
Crystal structure of the catalytic portion of human HMG-CoA reductase: insights into regulation of activity and catalysis.
Istvan, E.S., Palnitkar, M., Buchanan, S.K., Deisenhofer, J.(2000) EMBO J 19: 819-830
- PubMed: 10698924 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/19.5.819
- Primary Citation Related Structures:
1DQ8, 1DQ9, 1DQA - PubMed Abstract:
3-hydroxy-3-methylglutaryl-CoA reductase (HMGR) catalyzes the formation of mevalonate, the committed step in the biosynthesis of sterols and isoprenoids. The activity of HMGR is controlled through synthesis, degradation and phosphorylation to maintain the concentration of mevalonate-derived products. In addition to the physiological regulation of HMGR, the human enzyme has been targeted successfully by drugs in the clinical treatment of high serum cholesterol levels. Three crystal structures of the catalytic portion of human HMGR in complexes with HMG-CoA, with HMG and CoA, and with HMG, CoA and NADP(+), provide a detailed view of the enzyme active site. Catalytic portions of human HMGR form tight tetramers. The crystal structure explains the influence of the enzyme's oligomeric state on the activity and suggests a mechanism for cholesterol sensing. The active site architecture of human HMGR is different from that of bacterial HMGR; this may explain why binding of HMGR inhibitors to bacterial HMGRs has not been reported.
- Howard Hughes Medical Institute, Department of Biochemistry, University of Texas Southwestern Medical Center at Dallas, TX 75235-9050, USA.
Organizational Affiliation: