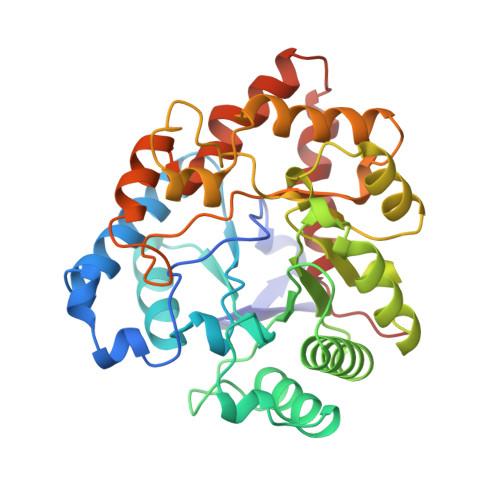
Three-dimensional structure of the zinc-containing phosphotriesterase with the bound substrate analog diethyl 4-methylbenzylphosphonate.
Vanhooke, J.L., Benning, M.M., Raushel, F.M., Holden, H.M.(1996) Biochemistry 35: 6020-6025
- PubMed: 8634243
- DOI: https://doi.org/10.1021/bi960325l
- Primary Citation Related Structures:
1DPM - PubMed Abstract:
Phosphotriesterase from Pseudomonas diminuta catalyzes the hydrolysis of paraoxon and related acetylcholinesterase inhibitors with rate enhancements that approach 10(12). The enzyme requires a binuclear metal center for activity and as isolated contains 2 equiv of zinc per subunit. Here we describe the three-dimensional structure of the Zn2+/Zn2+-substituted enzyme complexed with the substrate analog diethyl 4-methylbenzylphosphonate. Crystals employed in the investigation belonged to the space group C2 with unit cell dimensions of a = 129.6 A, b = 91.4 A, c = 69.4 A, beta = 91.9 degrees, and two subunits in the asymmetric unit. The model was refined by least-squares analysis to a nominal resolution of 2.1 A and a crystallographic R-factor of 15.4% for all measured X-ray data. As in the previously reported structure of the cadmium-containing enzyme, the bridging ligands are a carbamylated lysine residue (Lys 169) and a hydroxide. The zinc ions are separated by 3.3 A. The more buried zinc ion is surrounded by His 55, His 57, Lys 169, Asp 301, and the bridging hydroxide in a trigonal bipyramidal arrangement as described for the cadmium-substituted enzyme. Unlike the octahedral coordination observed for the more solvent-exposed cadmium ion, however, the second zinc is tetrahedrally ligated to Lys 169, His 201, His 230, and the bridging hydroxide. The diethyl 4-methylbenzylphosphonate occupies a site near the binuclear metal center with the phosphoryl oxygen of the substrate analog situated at 3.5 A from the more solvent-exposed zinc ion. The aromatic portion of the inhibitor binds in a fairly hydrophobic pocket. A striking feature of the active site pocket is the lack of direct electrostatic interactions between the inhibitor and the protein. This most likely explains the broad substrate specificity exhibited by phosphotriesterase. The position of the inhibitor within the active site suggests that the nucleophile for the hydrolysis reaction is the metal-bound hydroxide.
- Department of Biochemistry, University of Wisconsin, Madison 53705, USA.
Organizational Affiliation: