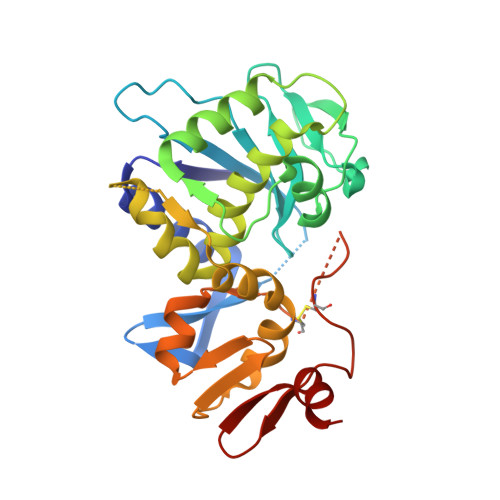
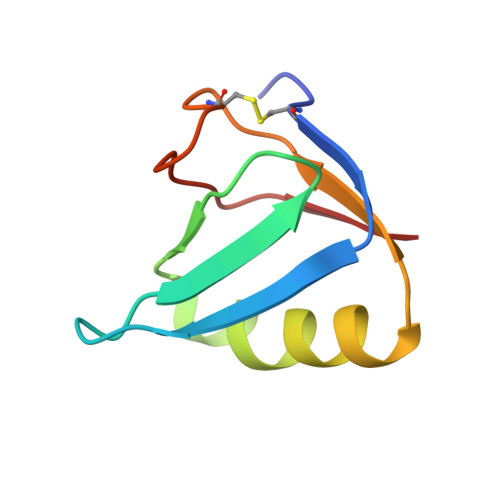
Crystal structure of the holotoxin from Shigella dysenteriae at 2.5 A resolution.
Fraser, M.E., Chernaia, M.M., Kozlov, Y.V., James, M.N.(1994) Nat Struct Biol 1: 59-64
- PubMed: 7656009 Search on PubMed
- DOI: https://doi.org/10.1038/nsb0194-59
- Primary Citation Related Structures:
1DM0 - PubMed Abstract:
Shigella dysenteriae is the pathogen responsible for the severe form of dysentery in humans. It produces Shiga toxin, the prototype of a family of closely related bacterial protein toxins. We have determined the structure of the holotoxin, an AB5 hexamer, by X-ray crystallography. The five B subunits form a pentameric ring, encircling a helix at the carboxy terminus of the A subunit. The A subunit interacts with the B pentamer via this C-terminal helix and a four-stranded mixed beta-sheet. The fold of the rest of the A subunit is similar to that of the A chain of the plant toxin ricin; both are N-glycosidases. However, the active site in the bacterial holotoxin is blocked by a segment of polypeptide chain. These residues of the A subunit would be released as part of the activation mechanism of the toxin.
- Department of Biochemistry, University of Alberta, Edmonton, Canada.
Organizational Affiliation: