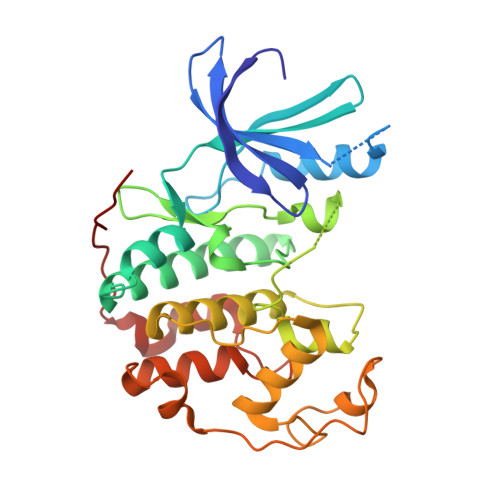
Binding mode of the 4-anilinoquinazoline class of protein kinase inhibitor: X-ray crystallographic studies of 4-anilinoquinazolines bound to cyclin-dependent kinase 2 and p38 kinase.
Shewchuk, L., Hassell, A., Wisely, B., Rocque, W., Holmes, W., Veal, J., Kuyper, L.F.(2000) J Med Chem 43: 133-138
- PubMed: 10633045 Search on PubMed
- DOI: https://doi.org/10.1021/jm990401t
- Primary Citation Related Structures:
1DI8, 1DI9 - PubMed Abstract:
4-Anilinoquinazolines represent an important class of protein kinase inhibitor. Modes of binding for two members of this inhibitor class were determined by X-ray crystallographic analysis of one inhibitor (4-[3-hydroxyanilino]-6,7-dimethoxyquinazoline) in complex with cyclin-dependent kinase 2 (CDK2) and the other (4-[3-methylsulfanylanilino]-6,7-dimethoxyquinazoline) in complex with p38 kinase. In both inhibitor/kinase structures, the 4-anilinoquinazoline was bound in the ATP site with the quinazoline ring system oriented along the peptide strand that links the two domains of the protein and with the anilino substituent projecting into a hydrophobic pocket within the protein interior. In each case, the nitrogen at position-1 of the quinazoline accepted a hydrogen bond from a backbone NH (CDK2, Leu-83; p38, Met-109) of the domain connector strand, and aromatic hydrogen atoms at C2 and C8 interacted with backbone carbonyl oxygen atoms of the peptide strand. The anilino group of the CDK2-bound compound was essentially coplanar with the quinazoline ring system and occupied a pocket between Lys-33 and Phe-80. For the p38-bound inhibitor, the anilino group was angled out of plane and was positioned between Lys-53 and Thr-106 in a manner similar to that observed for the aryl substituent of the pyridinylimidazole class of inhibitor.
- Glaxo Wellcome Inc., Five Moore Drive, Research Triangle Park, North Carolina 27709, USA.
Organizational Affiliation: