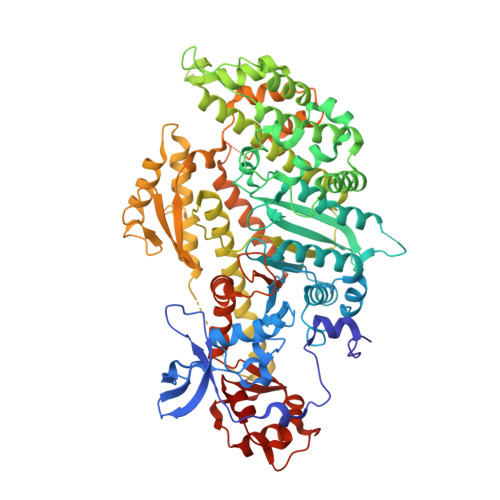
X-ray structures of the Dictyostelium discoideum myosin motor domain with six non-nucleotide analogs.
Gulick, A.M., Bauer, C.B., Thoden, J.B., Pate, E., Yount, R.G., Rayment, I.(2000) J Biological Chem 275: 398-408
- PubMed: 10617631 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.275.1.398
- Primary Citation Related Structures:
1D0X, 1D0Y, 1D0Z, 1D1A, 1D1B, 1D1C - PubMed Abstract:
The three-dimensional structures of the truncated myosin head from Dictyostelium discoideum myosin II complexed with dinitrophenylaminoethyl-, dinitrophenylaminopropyl-, o-nitrophenylaminoethyl-, m-nitrophenylaminoethyl-, p-nitrophenylaminoethyl-, and o-nitrophenyl-N-methyl-aminoethyl-diphosphate.beryllium fluoride have been determined to better than 2.3-A resolution. The structure of the protein and nucleotide binding pocket in these complexes is very similar to that of S1dC.ADP.BeF(x) (Fisher, A. J., Smith, C. A., Thoden, J., Smith, R., Sutoh, K., Holden, H. M., and Rayment, I. (1995) Biochemistry 34, 8960-8972). The position of the triphosphate-like moiety is essentially identical in all complexes. Furthermore, the alkyl-amino group plays the same role as the ribose by linking the triphosphate to the adenine binding pocket; however, none of the phenyl groups lie in the same position as adenine in S1dC.MgADP.BeF(x), even though several of these nucleotide analogs are functionally equivalent to ATP. Rather the former location of adenine is occupied by water in the nanolog complexes, and the phenyl groups are organized in a manner that attempts to optimize their hydrogen bonding interactions with this constellation of solvent molecules. A comparison of the kinetic and structural properties of the nanologs relative to ATP suggests that the ability of a substrate to sustain tension and to generate movement correlates with a well defined interaction with the active site water structure observed in S1dC.MgADP.BeF(x).
- Institute for Enzyme Research, Department of Biochemistry, University of Wisconsin, Madison, Wisconsin 53705, USA.
Organizational Affiliation: