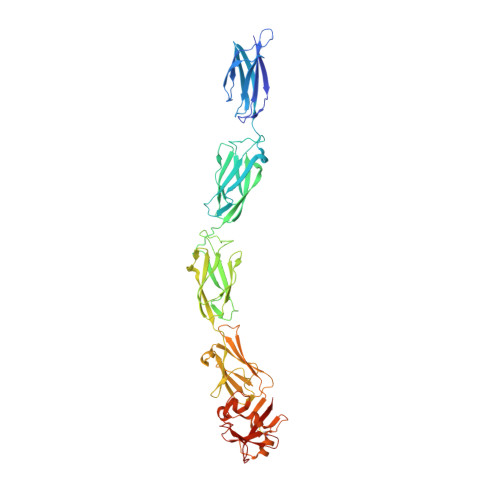Crystal structure of invasin: a bacterial integrin-binding protein.
Hamburger, Z.A., Brown, M.S., Isberg, R.R., Bjorkman, P.J.(1999) Science 286: 291-295
- PubMed: 10514372 Search on PubMed
- DOI: https://doi.org/10.1126/science.286.5438.291
- Primary Citation Related Structures:
1CWV - PubMed Abstract:
The Yersinia pseudotuberculosis invasin protein promotes bacterial entry by binding to host cell integrins with higher affinity than natural substrates such as fibronectin. The 2.3 angstrom crystal structure of the invasin extracellular region reveals five domains that form a 180 angstrom rod with structural similarities to tandem fibronectin type III domains. The integrin-binding surfaces of invasin and fibronectin include similarly located key residues, but in the context of different folds and surface shapes. The structures of invasin and fibronectin provide an example of convergent evolution, in which invasin presents an optimized surface for integrin binding, in comparison with host substrates.
- Division of Biology 156-29, Howard Hughes Medical Institute, California Institute of Technology, Pasadena, CA 91125, USA.
Organizational Affiliation: