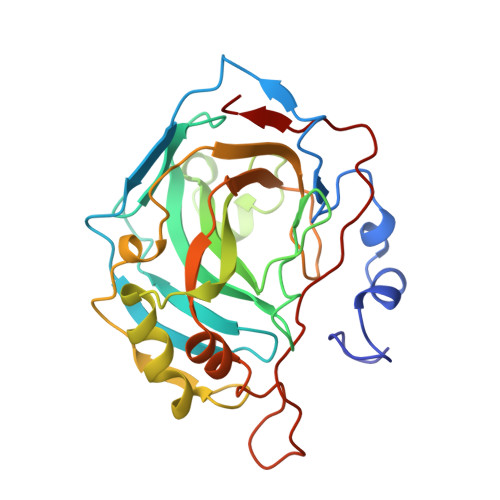
Crystal structure of the complex between human carbonic anhydrase II and the aromatic inhibitor 1,2,4-triazole.
Mangani, S., Liljas, A.(1993) J Mol Biology 232: 9-14
- PubMed: 8331673 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1993.1365
- Primary Citation Related Structures:
1CRA - PubMed Abstract:
The X-ray crystal structure of the complex between human carbonic anhydrase II and the inhibitor 1,2,4-triazole has been refined at 1.9 A resolution to a final R-factor of 0.153. Triazole is an analogue of the competitive inhibitor imidazole, but the crystal structure shows a different type of binding to the enzyme. 1,2,4-Triazole is directly bound to the zinc(II) ion through the nitrogen in position 4, replacing the native water/hydroxyl (Wat263) in a distorted four-co-ordinated complex. The interaction of the inhibitor with the active site is completed by two hydrogen bonds to O gamma of Thr200 and to the amide nitrogen atom of Thr199 through the two adjacent N-1 and N-2 atoms. The binding site of triazole overlaps the proposed binding sites for the substrates, explaining the observed competitive behaviour of the inhibitor towards CO2/HCO3- under equilibrium conditions.
- Department of Chemistry, University of Siena, Italy.
Organizational Affiliation: