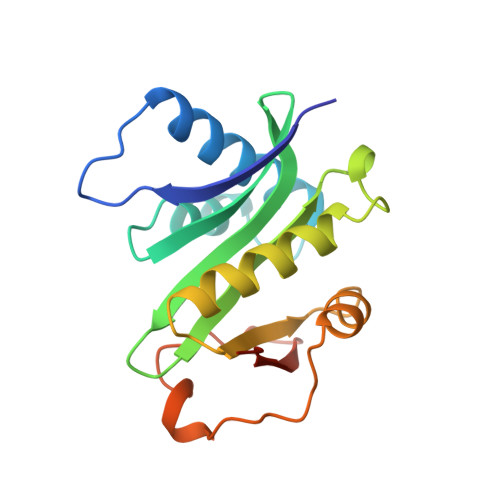
Crystal structure of the histone acetyltransferase domain of the human PCAF transcriptional regulator bound to coenzyme A.
Clements, A., Rojas, J.R., Trievel, R.C., Wang, L., Berger, S.L., Marmorstein, R.(1999) EMBO J 18: 3521-3532
- PubMed: 10393169 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/18.13.3521
- Primary Citation Related Structures:
1CM0 - PubMed Abstract:
The human p300/CBP-associating factor, PCAF, mediates transcriptional activation through its ability to acetylate nucleosomal histone substrates as well as transcriptional activators such as p53. We have determined the 2.3 A crystal structure of the histone acetyltransferase (HAT) domain of PCAF bound to coenzyme A. The structure reveals a central protein core associated with coenzyme A binding and a pronounced cleft that sits over the protein core and is flanked on opposite sides by the N- and C-terminal protein segments. A correlation of the structure with the extensive mutagenesis data for PCAF and the homologous yeast GCN5 protein implicates the cleft and the N- and C-terminal protein segments as playing an important role in histone substrate binding, and a glutamate residue in the protein core as playing an essential catalytic role. A structural comparison with the coenzyme-bound forms of the related N-acetyltransferases, HAT1 (yeast histone acetyltransferase 1) and SmAAT (Serratia marcescens aminoglycoside 3-N-acetyltransferase), suggests the mode of substrate binding and catalysis by these enzymes and establishes a paradigm for understanding the structure-function relationships of other enzymes that acetylate histones and transcriptional regulators to promote activated transcription.
- The Wistar Institute, University of Pennsylvania, Philadelphia, PA 19104, USA.
Organizational Affiliation: