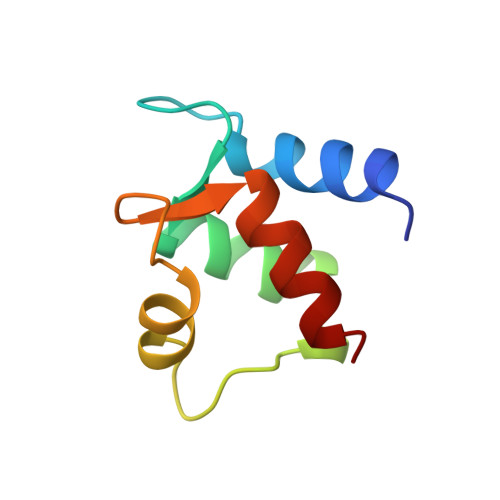
Determination of the solution structure of Apo calbindin D9k by NMR spectroscopy.
Skelton, N.J., Kordel, J., Chazin, W.J.(1995) J Mol Biology 249: 441-462
- PubMed: 7783203
- DOI: https://doi.org/10.1006/jmbi.1995.0308
- Primary Citation of Related Structures:
1CLB - PubMed Abstract:
The three-dimensional structure of apo calbindin D9k has been determined using constraints generated from nuclear magnetic resonance spectroscopy. The family of solution structures was calculated using a combination of distance geometry, restrained molecular dynamics, and hybrid relaxation matrix analysis of the nuclear Overhauser effect (NOE) cross-peak intensities. Errors and inconsistencies in the input constraints were identified using complete relaxation matrix analyses based on the results of preliminary structure calculations. The final input data consisted of 994 NOE distance constraints and 122 dihedral constraints, aided by the stereospecific assignment of the resonances from 21 beta-methylene groups and seven isopropyl groups of leucine and valine residues. The resulting family of 33 structures contain no violation of the distance constraints greater than 0.17 A or of the dihedral angle constraints greater than 10 degrees. The structures consist of a well-defined, antiparallel four-helix bundle, with a short anti-parallel beta-interaction between the two unoccupied calcium-binding loops. The root-mean-square deviation from the mean structure of the backbone heavy-atoms for the well-defined helical residues is 0.55 A. The remainder of the ion-binding loops, the linker loop connecting the two sub-domains of the protein, and the N and C termini exhibit considerable disorder between different structures in the ensemble. A comparison with the structure of the (Ca2+)2 state indicates that the largest changes associated with ion-binding occur in the middle of helix IV and in the packing of helix III onto the remainder of the protein. The change in conformation of these helices is associated with a subtle reorganization of many residues in the hydrophobic core, including some side-chains that are up to 15 A from the ion-binding site.
- Department of Protein Engineering, Genentech, Inc., South San Francisco, CA 94080, USA.
Organizational Affiliation: