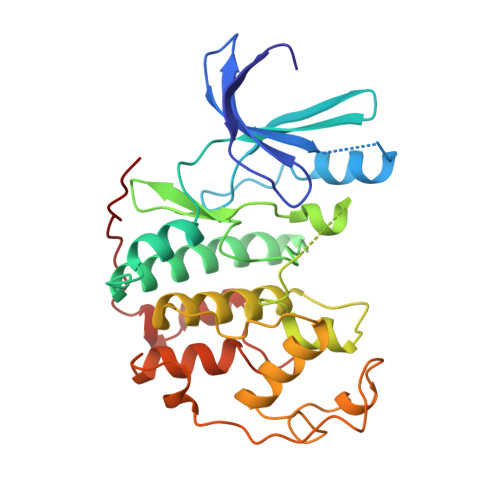
Exploiting chemical libraries, structure, and genomics in the search for kinase inhibitors.
Gray, N.S., Wodicka, L., Thunnissen, A.M., Norman, T.C., Kwon, S., Espinoza, F.H., Morgan, D.O., Barnes, G., LeClerc, S., Meijer, L., Kim, S.H., Lockhart, D.J., Schultz, P.G.(1998) Science 281: 533-538
- PubMed: 9677190 Search on PubMed
- DOI: https://doi.org/10.1126/science.281.5376.533
- Primary Citation Related Structures:
1CKP - PubMed Abstract:
Selective protein kinase inhibitors were developed on the basis of the unexpected binding mode of 2,6,9-trisubstituted purines to the adenosine triphosphate-binding site of the human cyclin-dependent kinase 2 (CDK2). By iterating chemical library synthesis and biological screening, potent inhibitors of the human CDK2-cyclin A kinase complex and of Saccharomyces cerevisiae Cdc28p were identified. The structural basis for the binding affinity and selectivity was determined by analysis of a three-dimensional crystal structure of a CDK2-inhibitor complex. The cellular effects of these compounds were characterized in mammalian cells and yeast. In the latter case the effects were characterized on a genome-wide scale by monitoring changes in messenger RNA levels in treated cells with high-density oligonucleotide probe arrays. Purine libraries could provide useful tools for analyzing a variety of signaling and regulatory pathways and may lead to the development of new therapeutics.
- Howard Hughes Medical Institute, University of California, Berkeley, CA 94720, USA.
Organizational Affiliation: