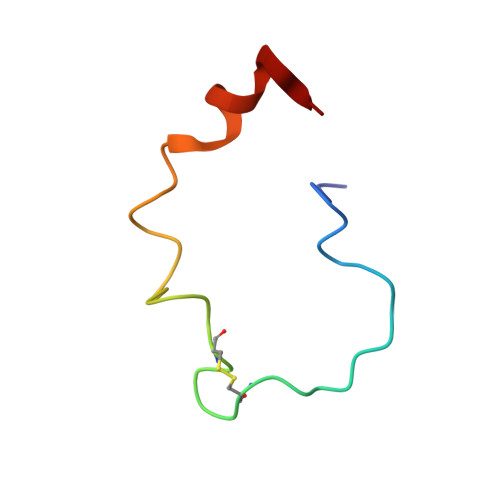
Structure of the metal-free gamma-carboxyglutamic acid-rich membrane binding region of factor IX by two-dimensional NMR spectroscopy.
Freedman, S.J., Furie, B.C., Furie, B., Baleja, J.D.(1995) J Biological Chem 270: 7980-7987
- PubMed: 7713897 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.270.14.7980
- Primary Citation Related Structures:
1CFH - PubMed Abstract:
The gamma-carboxyglutamic acid-rich domain of blood coagulation Factor IX is required for the binding of the protein to phospholipid membranes. To investigate the three-dimensional structure of this domain, a synthetic peptide corresponding to residues 1-47 of Factor IX was studied by 1H NMR spectroscopy. In the absence of metal ions, the proton chemical shift dispersion in the one-dimensional NMR spectrum indicated that the peptide contains regular structural elements. Upon the addition of Ca(II) or Mg(II), large chemical shift changes were observed in the amide proton and methyl proton regions of the spectrum, consistent with the conformational transitions that metal ions are known to induce in native Factor IX. The apopeptide was studied by two-dimensional NMR spectroscopy at 500 MHz to determine its solution structure. Protons were assigned using total correlation spectroscopy, nuclear Overhauser effect spectroscopy, and double quantum-filtered correlation spectroscopy experiments. Intensities of cross-peaks in the nuclear Overhauser effect spectrum were used to generate a set of interproton distance restraints. The structure of the apopeptide was then calculated using distance geometry methods. There are three structural elements in the apopeptide that are linked by a flexible polypeptide backbone. These elements include a short amino-terminal tetrapeptide loop (amino acids 6-9), the disulfide-containing hexapeptide loop (amino acids 18-23), and a carboxyl-terminal alpha helix (amino acids 37-46). Amide hydrogen exchange kinetics indicate that the majority of the peptide is solvent accessible, except in the carboxyl-terminal element. The structured regions in the apopeptide are insufficient to support phospholipid binding, indicating the importance of additional structural features in the Ca(II)-stabilized conformer.
- Center for Hemostasis and Thrombosis Research, New England Medical Center, Boston, Massachusetts, USA.
Organizational Affiliation: