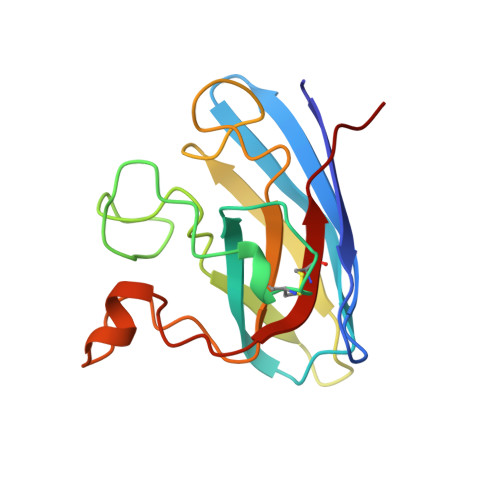
Crystallographic structures of bovine copper-zinc superoxide dismutase reveal asymmetry in two subunits: functionally important three and five coordinate copper sites captured in the same crystal.
Hough, M.A., Hasnain, S.S.(1999) J Mol Biology 287: 579-592
- PubMed: 10092461 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1999.2610
- Primary Citation Related Structures:
1CB4, 1CBJ - PubMed Abstract:
A key feature of the generally accepted catalytic mechanism of CuZn superoxide dismutase (CuZnSOD) is the breakage of the imidazolate bridge between copper and zinc and the loss of a coordinated water molecule from copper on reduction from Cu(II) to Cu(I). Crystal structures exist for the enzyme from a number of sources in the oxidised, five coordinate copper form. For the reduced form two structures from different sources have been determined only recently but provide contradictory results. We present crystal structures of bovine CuZnSOD (BSOD) in two different space groups. The structure of the P212121 form (pBSOD), at 1.65 A resolution clearly shows one subunit with Cu in the five coordinate, oxidised form, and the other with Cu in the three coordinate form expected for the reduced state. This mixed state of pBSOD is confirmed by XANES data of these crystals. The pBSOD structure has thus captured each subunit in one of the two oxidation state conformations and thus provides direct crystallographic evidence for the superoxide dismutase mechanism involving the breakage of the imidazole bridge between Cu and Zn. A shift in the position of copper in subunit A poises the catalytic centre to undergo the first stage of catalysis via dissociation of Cu from His61 with a concomittant movement of the coordinated water molecule towards His61, which rotates by approximately 20 degrees, enabling it to form a hydrogen bond to the water molecule. The Cu-Zn separation in the reduced site is increased by approximately 0.5 A. In contrast the 2.3 A resolution structure in space group C2221 (cBSOD) shows both of the Cu atoms to be in the five coordinate, oxidised form but in this space group the whole of subunit A is significantly more disordered than subunit B. An examination of published structures of "oxidised" SODs, shows a trend towards longer Cu-Zn and Cu-His61 separations in subunit A, which together with the structures reported here indicate a potential functional asymmetry between the subunits of CuZnSODs. We also suggest that the increased separation between Cu and Zn is a precursor to breakage of His61.
- CLRC Daresbury Laboratory, Daresbury, Warrington, Cheshire, WA4 4AD, UK.
Organizational Affiliation: