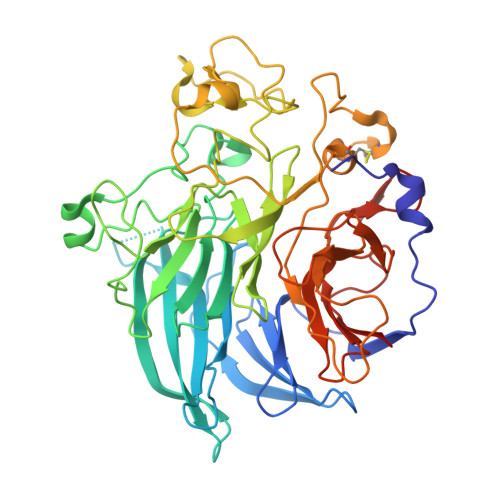
Structure and mechanism of soluble quinoprotein glucose dehydrogenase.
Oubrie, A., Rozeboom, H.J., Kalk, K.H., Olsthoorn, A.J., Duine, J.A., Dijkstra, B.W.(1999) EMBO J 18: 5187-5194
- PubMed: 10508152 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/18.19.5187
- Primary Citation Related Structures:
1C9U, 1CQ1 - PubMed Abstract:
Soluble glucose dehydrogenase (s-GDH; EC 1.1.99.17) is a classical quinoprotein which requires the cofactor pyrroloquinoline quinone (PQQ) to oxidize glucose to gluconolactone. The reaction mechanism of PQQ-dependent enzymes has remained controversial due to the absence of comprehensive structural data. We have determined the X-ray structure of s-GDH with the cofactor at 2.2 A resolution, and of a complex with reduced PQQ and glucose at 1.9 A resolution. These structures reveal the active site of s-GDH, and show for the first time how a functionally bound substrate interacts with the cofactor in a PQQ-dependent enzyme. Twenty years after the discovery of PQQ, our results finally provide conclusive evidence for a reaction mechanism comprising general base-catalyzed hydride transfer, rather than the generally accepted covalent addition-elimination mechanism. Thus, PQQ-dependent enzymes use a mechanism similar to that of nicotinamide- and flavin-dependent oxidoreductases.
- Laboratory of Biophysical Chemistry and BIOSON Research Institute, University of Groningen, Nijenborgh 4, 9747 AG Groningen.
Organizational Affiliation: